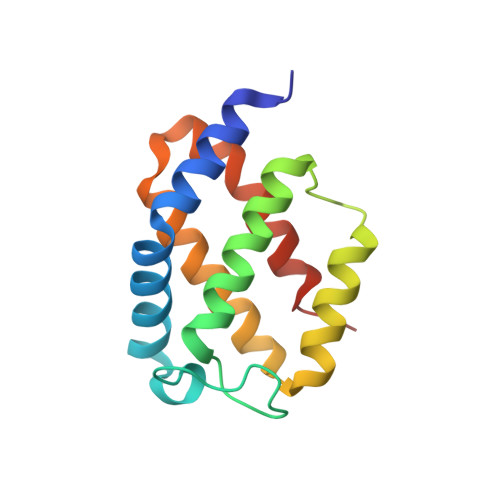
Crystal structure of a biliverdin-bound phycobiliprotein: Interdependence of oligomerization and chromophorylation.
Fuenzalida-Werner, J.P., Janowski, R., Mishra, K., Weidenfeld, I., Niessing, D., Ntziachristos, V., Stiel, A.C.(2018) J Struct Biol 204: 519-522
- PubMed: 30287387
- DOI: https://doi.org/10.1016/j.jsb.2018.09.013
- Primary Citation Related Structures:
6FZN, 6FZO - PubMed Abstract:
Small, ultra-red fluorescence protein (smURFP) introduces the non-native biliverdin (BV) chromophore to phycobiliproteins (PBPs), allowing them to be used as transgenic labels for in vivo mammalian imaging. Presently, no structural information exists for PBPs bound to the non-native BV chromophore, which limits the further development of smURFP and related proteins as imaging labels or indicators. Here we describe the first crystal structure of a PBP bound to BV. The structures of smURFP-Y56R with BV and smURFP-Y56F without BV reveal unique oligomerization interfaces different from those in wild-type PBPs bound to native chromophores. Our structures suggest that the oligomerization interface affects the BV binding site, creating a link between oligomerization and chromophorylation that we confirmed through site-directed mutagenesis and that may help guide efforts to improve the notorious chromophorylation of smURFP and other PBPs engineered to bind BV.
- Institute of Biological and Medical Imaging (IBMI), Helmholtz Zentrum München, Neuherberg, Germany.
Organizational Affiliation: