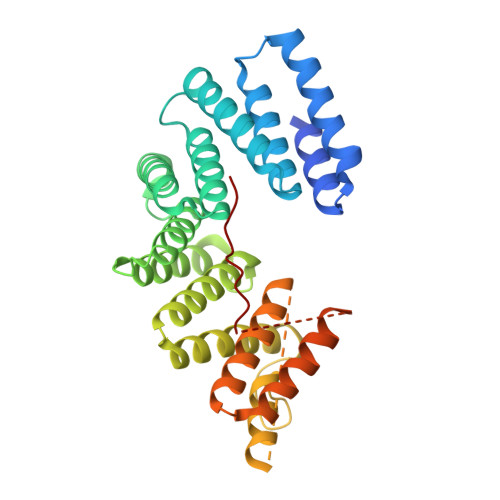
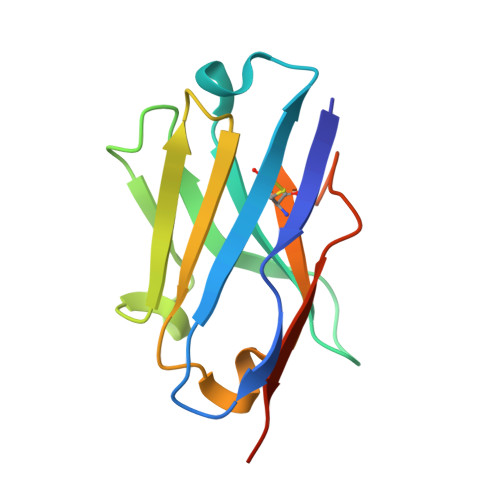
Structural basis for isoform-specific kinesin-1 recognition of Y-acidic cargo adaptors.
Pernigo, S., Chegkazi, M.S., Yip, Y.Y., Treacy, C., Glorani, G., Hansen, K., Politis, A., Bui, S., Dodding, M.P., Steiner, R.A.(2018) Elife 7
- PubMed: 30320553 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.38362
- Primary Citation Related Structures:
6FUZ, 6FV0 - PubMed Abstract:
The light chains (KLCs) of the heterotetrameric microtubule motor kinesin-1, that bind to cargo adaptor proteins and regulate its activity, have a capacity to recognize short peptides via their tetratricopeptide repeat domains (KLC TPR ). Here, using X-ray crystallography, we show how kinesin-1 recognizes a novel class of adaptor motifs that we call 'Y-acidic' (tyrosine flanked by acidic residues), in a KLC-isoform-specific manner. Binding specificities of Y-acidic motifs (present in JIP1 and in TorsinA) to KLC1 TPR are distinct from those utilized for the recognition of W-acidic motifs, found in adaptors, that are KLC-isoform non-selective. However, a partial overlap on their receptor-binding sites implies that adaptors relying on Y-acidic and W-acidic motifs must act independently. We propose a model to explain why these two classes of motifs that bind to the concave surface of KLC TPR with similar low micromolar affinity can exhibit different capacities to promote kinesin-1 activity.
- Randall Centre of Cell and Molecular Biophysics, Faculty of Life Sciences and Medicine, King's College London, London, United Kingdom.
Organizational Affiliation: