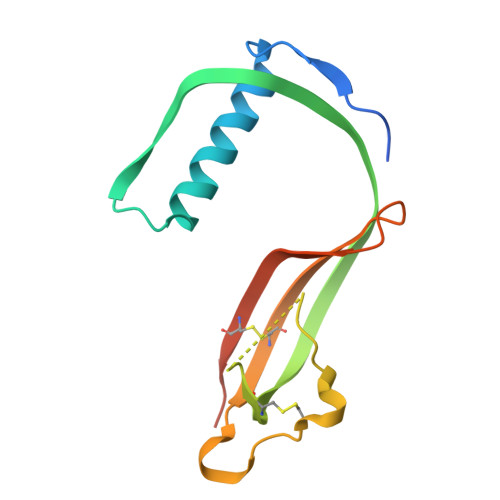
Structural and functional analysis of cystatin E reveals enzymologically relevant dimer and amyloid fibril states.
Dall, E., Hollerweger, J.C., Dahms, S.O., Cui, H., Haussermann, K., Brandstetter, H.(2018) J Biological Chem 293: 13151-13165
- PubMed: 29967063
- DOI: https://doi.org/10.1074/jbc.RA118.002154
- Primary Citation of Related Structures:
6FK0 - PubMed Abstract:
Protein activity is often regulated by altering the oligomerization state. One mechanism of multimerization involves domain swapping, wherein proteins exchange parts of their structures and thereby form long-lived dimers or multimers. Domain swapping has been specifically observed in amyloidogenic proteins, for example the cystatin superfamily of cysteine protease inhibitors. Cystatins are twin-headed inhibitors, simultaneously targeting the lysosomal cathepsins and legumain, with important roles in cancer progression and Alzheimer's disease. Although cystatin E is the most potent legumain inhibitor identified so far, nothing is known about its propensity to oligomerize. In this study, we show that conformational destabilization of cystatin E leads to the formation of a domain-swapped dimer with increased conformational stability. This dimer was active as a legumain inhibitor by forming a trimeric complex. By contrast, the binding sites toward papain-like proteases were buried within the cystatin E dimer. We also showed that the dimers could further convert to amyloid fibrils. Unexpectedly, cystatin E amyloid fibrils contained functional protein, which inhibited both legumain and papain-like enzymes. Fibril formation was further regulated by glycosylation. We speculate that cystatin amyloid fibrils might serve as a binding platform to stabilize the pH-sensitive legumain and cathepsins in the extracellular environment, contributing to their physiological and pathological functions.
- From the Department of Biosciences, University of Salzburg, A-5020 Salzburg, Austria and.
Organizational Affiliation: