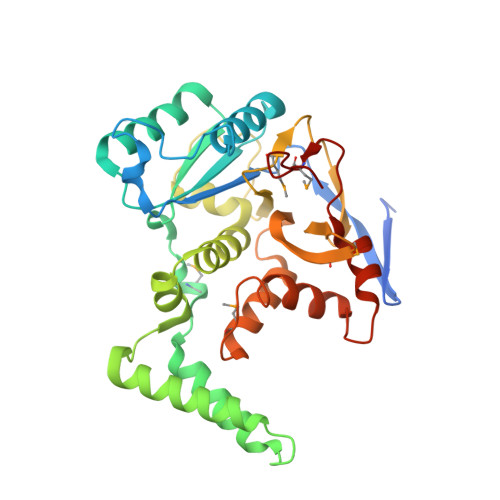
Structural basis for the glycosyltransferase activity of theSalmonellaeffector SseK3.
Esposito, D., Gunster, R.A., Martino, L., El Omari, K., Wagner, A., Thurston, T.L.M., Rittinger, K.(2018) J Biological Chem 293: 5064-5078
- PubMed: 29449376
- DOI: https://doi.org/10.1074/jbc.RA118.001796
- Primary Citation Related Structures:
6EYR, 6EYT - PubMed Abstract:
The Salmonella -secreted effector SseK3 translocates into host cells, targeting innate immune responses, including NF-κB activation. SseK3 is a glycosyltransferase that transfers an N -acetylglucosamine (GlcNAc) moiety onto the guanidino group of a target arginine, modulating host cell function. However, a lack of structural information has precluded elucidation of the molecular mechanisms in arginine and GlcNAc selection. We report here the crystal structure of SseK3 in its apo form and in complex with hydrolyzed UDP-GlcNAc. SseK3 possesses the typical glycosyltransferase type-A (GT-A)-family fold and the metal-coordinating D X D motif essential for ligand binding and enzymatic activity. Several conserved residues were essential for arginine GlcNAcylation and SseK3-mediated inhibition of NF-κB activation. Isothermal titration calorimetry revealed SseK3's preference for manganese coordination. The pattern of interactions in the substrate-bound SseK3 structure explained the selection of the primary ligand. Structural rearrangement of the C-terminal residues upon ligand binding was crucial for SseK3's catalytic activity, and NMR analysis indicated that SseK3 has limited UDP-GlcNAc hydrolysis activity. The release of free N -acetyl α-d-glucosamine, and the presence of the same molecule in the SseK3 active site, classified it as a retaining glycosyltransferase. A glutamate residue in the active site suggested a double-inversion mechanism for the arginine N -glycosylation reaction. Homology models of SseK1, SseK2, and the Escherichia coli orthologue NleB1 reveal differences in the surface electrostatic charge distribution, possibly accounting for their diverse activities. This first structure of a retaining GT-A arginine N -glycosyltransferase provides an important step toward a better understanding of this enzyme class and their roles as bacterial effectors.
- From the Molecular Structure of Cell Signalling Laboratory, Francis Crick Institute, 1 Midland Road, London NW1 1AT, United Kingdom.
Organizational Affiliation: