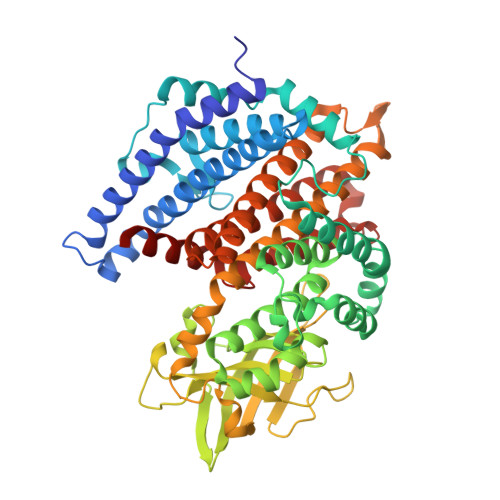
Structural basis for substrate specificity of methylsuccinyl-CoA dehydrogenase, an unusual member of the acyl-CoA dehydrogenase family.
Schwander, T., McLean, R., Zarzycki, J., Erb, T.J.(2018) J Biological Chem 293: 1702-1712
- PubMed: 29275330 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA117.000764
- Primary Citation Related Structures:
6ES9 - PubMed Abstract:
(2 S )-methylsuccinyl-CoA dehydrogenase (MCD) belongs to the family of FAD-dependent acyl-CoA dehydrogenase (ACD) and is a key enzyme of the ethylmalonyl-CoA pathway for acetate assimilation. It catalyzes the oxidation of (2 S )-methylsuccinyl-CoA to α,β-unsaturated mesaconyl-CoA and shows only about 0.5% activity with succinyl-CoA. Here we report the crystal structure of MCD at a resolution of 1.37 Å. The enzyme forms a homodimer of two 60-kDa subunits. Compared with other ACDs, MCD contains an ∼170-residue-long N-terminal extension that structurally mimics a dimer-dimer interface of these enzymes that are canonically organized as tetramers. MCD catalyzes the unprecedented oxidation of an α-methyl branched dicarboxylic acid CoA thioester. Substrate specificity is achieved by a cluster of three arginines that accommodates the terminal carboxyl group and a dedicated cavity that facilitates binding of the C2 methyl branch. MCD apparently evolved toward preventing the nonspecific oxidation of succinyl-CoA, which is a close structural homolog of (2 S )-methylsuccinyl-CoA and an essential intermediate in central carbon metabolism. For different metabolic engineering and biotechnological applications, however, an enzyme that can oxidize succinyl-CoA to fumaryl-CoA is sought after. Based on the MCD structure, we were able to shift substrate specificity of MCD toward succinyl-CoA through active-site mutagenesis.
- From the Department of Biochemistry and Synthetic Metabolism, Max-Planck-Institute for Terrestrial Microbiology Marburg, Karl-von-Frisch-Strasse 10, D-35043 Marburg, Germany and.
Organizational Affiliation: