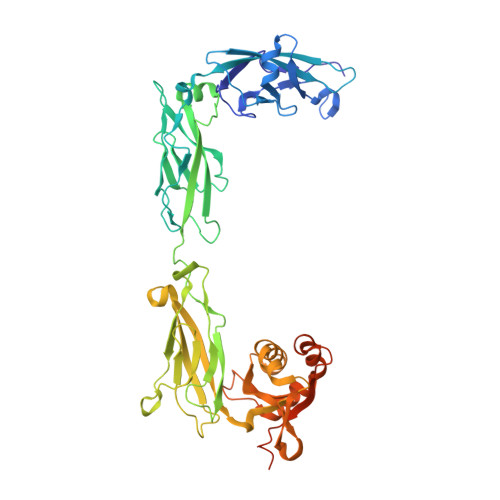Structural determinants of protocadherin-15 mechanics and function in hearing and balance perception.
Choudhary, D., Narui, Y., Neel, B.L., Wimalasena, L.N., Klanseck, C.F., De-la-Torre, P., Chen, C., Araya-Secchi, R., Tamilselvan, E., Sotomayor, M.(2020) Proc Natl Acad Sci U S A
- PubMed: 32963095
- DOI: https://doi.org/10.1073/pnas.1920444117
- Primary Citation of Related Structures:
5TPK, 5ULY, 5W1D, 6BWN, 6BXU, 6E8F, 6EB5, 6EET, 6MFO, 6N22, 6N2E - PubMed Abstract:
The vertebrate inner ear, responsible for hearing and balance, is able to sense minute mechanical stimuli originating from an extraordinarily broad range of sound frequencies and intensities or from head movements. Integral to these processes is the tip-link protein complex, which conveys force to open the inner-ear transduction channels that mediate sensory perception. Protocadherin-15 and cadherin-23, two atypically large cadherins with 11 and 27 extracellular cadherin (EC) repeats, are involved in deafness and balance disorders and assemble as parallel homodimers that interact to form the tip link. Here we report the X-ray crystal structure of a protocadherin-15 + cadherin-23 heterotetrameric complex at 2.9-Å resolution, depicting a parallel homodimer of protocadherin-15 EC1-3 molecules forming an antiparallel complex with two cadherin-23 EC1-2 molecules. In addition, we report structures for 10 protocadherin-15 fragments used to build complete high-resolution models of the monomeric protocadherin-15 ectodomain. Molecular dynamics simulations and validated crystal contacts are used to propose models for the complete extracellular protocadherin-15 parallel homodimer and the tip-link bond. Steered molecular dynamics simulations of these models suggest conditions in which a structurally diverse and multimodal protocadherin-15 ectodomain can act as a stiff or soft gating spring. These results reveal the structural determinants of tip-link-mediated inner-ear sensory perception and elucidate protocadherin-15's structural and adhesive properties relevant in disease.
- Department of Chemistry and Biochemistry, The Ohio State University, Columbus, OH 43210.
Organizational Affiliation: