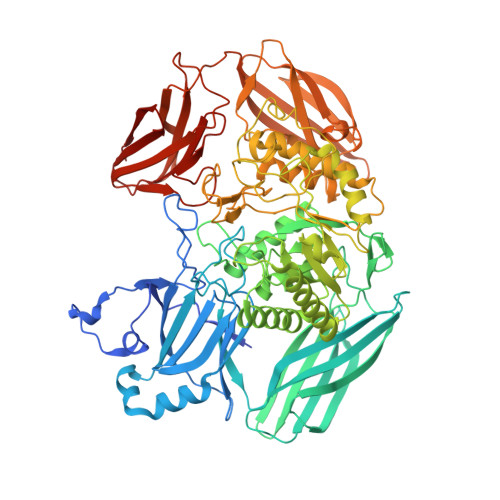
Active site flexibility revealed in crystal structures of Parabacteroides merdae beta-glucuronidase from the human gut microbiome.
Little, M.S., Ervin, S.M., Walton, W.G., Tripathy, A., Xu, Y., Liu, J., Redinbo, M.R.(2018) Protein Sci 27: 2010-2022
- PubMed: 30230652 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3507
- Primary Citation Related Structures:
6D7J, 6DXU - PubMed Abstract:
β-Glucuronidase (GUS) enzymes in the gastrointestinal tract are involved in maintaining mammalian-microbial symbiosis and can play key roles in drug efficacy and toxicity. Parabacteroides merdae GUS was identified as an abundant mini-Loop 2 (mL2) type GUS enzyme in the Human Microbiome Project gut metagenomic database. Here, we report the crystal structure of P. merdae GUS and highlight the differences between this enzyme and extant structures of gut microbial GUS proteins. We find that P. merdae GUS exhibits a distinct tetrameric quaternary structure and that the mL2 motif traces a unique path within the active site, which also includes two arginines distinctive to this GUS. We observe two states of the P. merdae GUS active site; a loop repositions itself by more than 50 Å to place a functionally-relevant residue into the enzyme's catalytic site. Finally, we find that P. merdae GUS is able to bind to homo and heteropolymers of the polysaccharide alginic acid. Together, these data broaden our understanding of the structural and functional diversity in the GUS family of enzymes present in the human gut microbiome and point to specialization as an important feature of microbial GUS orthologs.
- Department of Chemistry, University of North Carolina, Chapel Hill, North Carolina, 27599-3290.
Organizational Affiliation: