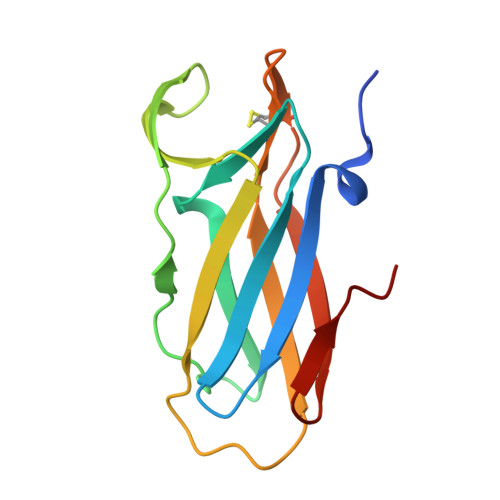
Structural and biochemical insights into the disulfide reductase mechanism of DsbD, an essential enzyme for neisserial pathogens.
Smith, R.P., Mohanty, B., Mowlaboccus, S., Paxman, J.J., Williams, M.L., Headey, S.J., Wang, G., Subedi, P., Doak, B.C., Kahler, C.M., Scanlon, M.J., Heras, B.(2018) J Biological Chem 293: 16559-16571
- PubMed: 30181210 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.004847
- Primary Citation Related Structures:
6DNL, 6DNU, 6DNV, 6DPS - PubMed Abstract:
The worldwide incidence of neisserial infections, particularly gonococcal infections, is increasingly associated with antibiotic-resistant strains. In particular, extensively drug-resistant Neisseria gonorrhoeae strains that are resistant to third-generation cephalosporins are a major public health concern. There is a pressing clinical need to identify new targets for the development of antibiotics effective against Neisseria -specific processes. In this study, we report that the bacterial disulfide reductase DsbD is highly prevalent and conserved among Neisseria spp. and that this enzyme is essential for survival of N. gonorrhoeae DsbD is a membrane-bound protein that consists of two periplasmic domains, n-DsbD and c-DsbD, which flank the transmembrane domain t-DsbD. In this work, we show that the two functionally essential periplasmic domains of Neisseria DsbD catalyze electron transfer reactions through unidirectional interdomain interactions, from reduced c-DsbD to oxidized n-DsbD, and that this process is not dictated by their redox potentials. Structural characterization of the Neisseria n- and c-DsbD domains in both redox states provides evidence that steric hindrance reduces interactions between the two periplasmic domains when n-DsbD is reduced, thereby preventing a futile redox cycle. Finally, we propose a conserved mechanism of electron transfer for DsbD and define the residues involved in domain-domain recognition. Inhibitors of the interaction of the two DsbD domains have the potential to be developed as anti-neisserial agents.
- From the Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne 3086, Victoria, Australia.
Organizational Affiliation: