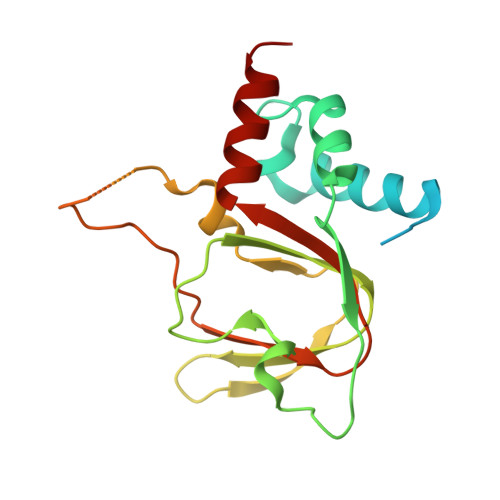
The cyclic nucleotide-binding homology domain of the integral membrane protein CNNM mediates dimerization and is required for Mg2+efflux activity.
Chen, Y.S., Kozlov, G., Fakih, R., Funato, Y., Miki, H., Gehring, K.(2018) J Biological Chem 293: 19998-20007
- PubMed: 30341174 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.005672
- Primary Citation Related Structures:
6DFD, 6DJ3 - PubMed Abstract:
Proteins of the cyclin M family (CNNMs; also called ancient conserved domain proteins, or ACDPs) are represented by four integral membrane proteins that have been proposed to function as Mg 2+ transporters. CNNMs are associated with a number of genetic diseases affecting ion movement and cancer via their association with highly oncogenic phosphatases of regenerating liver (PRLs). Structurally, CNNMs contain an N-terminal extracellular domain, a transmembrane domain (DUF21), and a large cytosolic region containing a cystathionine-β-synthase (CBS) domain and a putative cyclic nucleotide-binding homology (CNBH) domain. Although the CBS domain has been extensively characterized, little is known about the CNBH domain. Here, we determined the first crystal structures of the CNBH domains of CNNM2 and CNNM3 at 2.6 and 1.9 Å resolutions. Contrary to expectation, these domains did not bind cyclic nucleotides, but mediated dimerization both in crystals and in solution. Analytical ultracentrifugation experiments revealed an inverse correlation between the propensity of the CNBH domains to dimerize and the ability of CNNMs to mediate Mg 2+ efflux. CNBH domains from active family members were observed as both dimers and monomers, whereas the inactive member, CNNM3, was observed only as a dimer. Mutational analysis revealed that the CNBH domain was required for Mg 2+ efflux activity of CNNM4. This work provides a structural basis for understanding the function of CNNM proteins in Mg 2+ transport and associated diseases.
- From the Department of Biochemistry and Centre for Structural Biology, McGill University, Montreal, Quebec H3G 0B1, Canada and.
Organizational Affiliation: