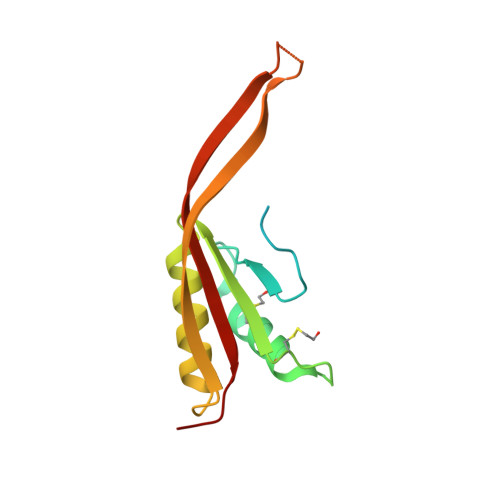
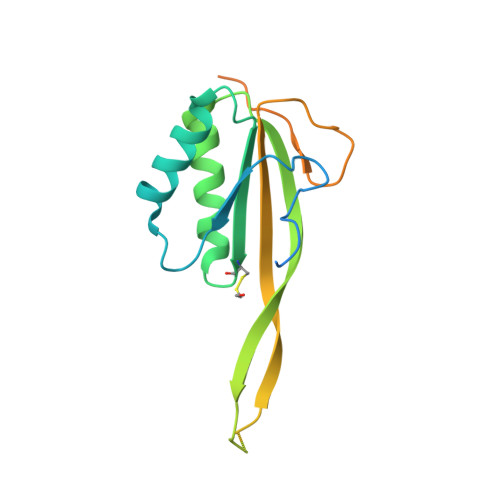
Crystal Structure of Human Rpp20/Rpp25 Reveals Quaternary Level Adaptation of the Alba Scaffold as Structural Basis for Single-stranded RNA Binding.
Chan, C.W., Kiesel, B.R., Mondragon, A.(2018) J Mol Biology 430: 1403-1416
- PubMed: 29625199 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2018.03.029
- Primary Citation Related Structures:
6CWX - PubMed Abstract:
Ribonuclease P (RNase P) catalyzes the removal of 5' leaders of tRNA precursors and its central catalytic RNA subunit is highly conserved across all domains of life. In eukaryotes, RNase P and RNase MRP, a closely related ribonucleoprotein enzyme, share several of the same protein subunits, contain a similar catalytic RNA core, and exhibit structural features that do not exist in their bacterial or archaeal counterparts. A unique feature of eukaryotic RNase P/MRP is the presence of two relatively long and unpaired internal loops within the P3 region of their RNA subunit bound by a heterodimeric protein complex, Rpp20/Rpp25. Here we present a crystal structure of the human Rpp20/Rpp25 heterodimer and we propose, using comparative structural analyses, that the evolutionary divergence of the single-stranded and helical nucleic acid binding specificities of eukaryotic Rpp20/Rpp25 and their related archaeal Alba chromatin protein dimers, respectively, originate primarily from quaternary level differences observed in their heterodimerization interface. Our work provides structural insights into how the archaeal Alba protein scaffold was adapted evolutionarily for incorporation into several functionally-independent eukaryotic ribonucleoprotein complexes.
- Department of Molecular Biosciences, Northwestern University, 2205 Tech Drive, Evanston, IL 60208-3500, United States.
Organizational Affiliation: