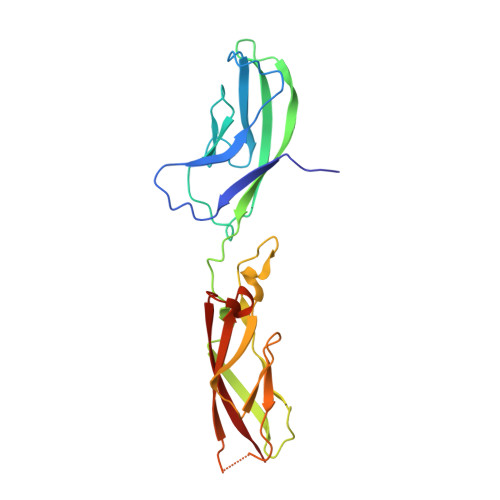
Homophilic and Heterophilic Interactions of Type II Cadherins Identify Specificity Groups Underlying Cell-Adhesive Behavior.
Brasch, J., Katsamba, P.S., Harrison, O.J., Ahlsen, G., Troyanovsky, R.B., Indra, I., Kaczynska, A., Kaeser, B., Troyanovsky, S., Honig, B., Shapiro, L.(2018) Cell Rep 23: 1840-1852
- PubMed: 29742438
- DOI: https://doi.org/10.1016/j.celrep.2018.04.012
- Primary Citation Related Structures:
6CG6, 6CG7, 6CGB, 6CGS, 6CGU - PubMed Abstract:
Type II cadherins are cell-cell adhesion proteins critical for tissue patterning and neuronal targeting but whose molecular binding code remains poorly understood. Here, we delineate binding preferences for type II cadherin cell-adhesive regions, revealing extensive heterophilic interactions between specific pairs, in addition to homophilic interactions. Three distinct specificity groups emerge from our analysis with members that share highly similar heterophilic binding patterns and favor binding to one another. Structures of adhesive fragments from each specificity group confirm near-identical dimer topology conserved throughout the family, allowing interface residues whose conservation corresponds to specificity preferences to be identified. We show that targeted mutation of these residues converts binding preferences between specificity groups in biophysical and co-culture assays. Our results provide a detailed understanding of the type II cadherin interaction map and a basis for defining their role in tissue patterning and for the emerging importance of their heterophilic interactions in neural connectivity.
- Zuckerman Mind Brain Behavior Institute, New York, NY 10027, USA; Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: