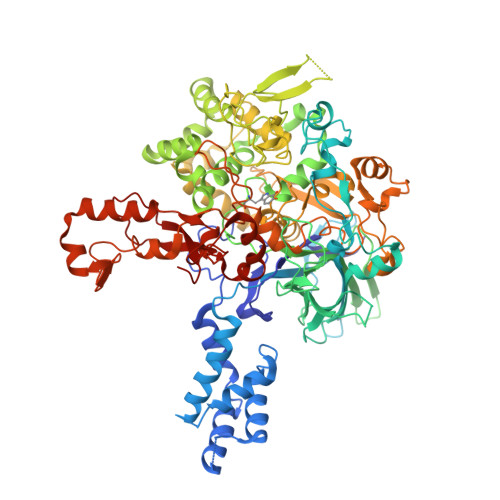
Structure and Enzymatic Properties of an Unusual Cysteine Tryptophylquinone-Dependent Glycine Oxidase from Pseudoalteromonas luteoviolacea.
Andreo-Vidal, A., Mamounis, K.J., Sehanobish, E., Avalos, D., Campillo-Brocal, J.C., Sanchez-Amat, A., Yukl, E.T., Davidson, V.L.(2018) Biochemistry 57: 1155-1165
- PubMed: 29381339
- DOI: https://doi.org/10.1021/acs.biochem.8b00009
- Primary Citation Related Structures:
6BYW - PubMed Abstract:
Glycine oxidase from Pseudoalteromonas luteoviolacea (PlGoxA) is a cysteine tryptophylquinone (CTQ)-dependent enzyme. Sequence analysis and phylogenetic analysis place it in a newly designated subgroup (group IID) of a recently identified family of LodA-like proteins, which are predicted to possess CTQ. The crystal structure of PlGoxA reveals that it is a homotetramer. It possesses an N-terminal domain with no close structural homologues in the Protein Data Bank. The active site is quite small because of intersubunit interactions, which may account for the observed cooperativy toward glycine. Steady-state kinetic analysis yielded the following values: k cat = 6.0 ± 0.2 s -1 , K 0.5 = 187 ± 18 μM, and h = 1.77 ± 0.27. In contrast to other quinoprotein amine dehydrogenases and oxidases that exhibit anomalously large primary kinetic isotope effects on the rate of reduction of the quinone cofactor by the amine substrate, no significant primary kinetic isotope effect was observed for this reaction of PlGoxA. The absorbance spectrum of glycine-reduced PlGoxA exhibits features in the range of 400-650 nm that have not previously been seen in other quinoproteins. Thus, in addition to the unusual structural features of PlGoxA, the kinetic and chemical reaction mechanisms of the reductive half-reaction of PlGoxA appear to be distinct from those of other amine dehydrogenases and amine oxidases that use tryptophylquinone and tyrosylquinone cofactors.
- Department of Genetics and Microbiology, University of Murcia , Murcia 30100, Spain.
Organizational Affiliation: