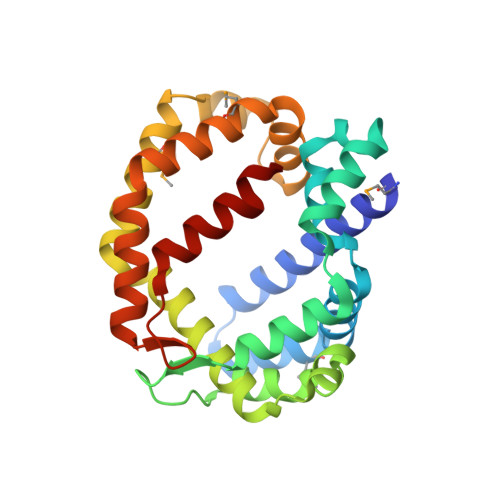
Structure of a lipid-bound viral membrane assembly protein reveals a modality for enclosing the lipid bilayer.
Pathak, P.K., Peng, S., Meng, X., Han, Y., Zhang, B., Zhang, F., Xiang, Y., Deng, J.(2018) Proc Natl Acad Sci U S A 115: 7028-7032
- PubMed: 29915071
- DOI: https://doi.org/10.1073/pnas.1805855115
- Primary Citation Related Structures:
6BR8, 6BR9, 6CB6, 6CB7 - PubMed Abstract:
Cellular membranes are maintained as closed compartments, broken up only transiently during membrane reorganization or lipid transportation. However, open-ended membranes, likely derived from scissions of the endoplasmic reticulum, persist in vaccinia virus-infected cells during the assembly of the viral envelope. A group of viral membrane assembly proteins (VMAPs) were identified as essential for this process. To understand the mechanism of VMAPs, we determined the 2.2-Å crystal structure of the largest member, named A6, which is a soluble protein with two distinct domains. The structure of A6 displays a novel protein fold composed mainly of alpha helices. The larger C-terminal domain forms a unique cage that encloses multiple glycerophospholipids with a lipid bilayer-like configuration. The smaller N-terminal domain does not bind lipid but negatively affects lipid binding by A6. Mutations of key hydrophobic residues lining the lipid-binding cage disrupt lipid binding and abolish viral replication. Our results reveal a protein modality for enclosing the lipid bilayer and provide molecular insight into a viral machinery involved in generating and/or stabilizing open-ended membranes.
- Department of Biochemistry and Molecular Biology, Oklahoma State University, Stillwater, OK 74078.
Organizational Affiliation: