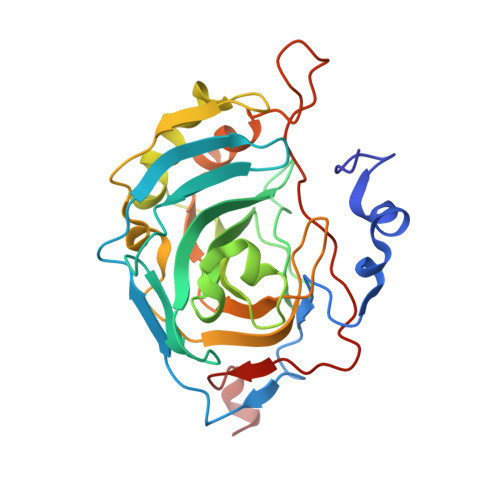
Structural insights into a thermostable variant of human carbonic anhydrase II.
Kean, K.M., Porter, J.J., Mehl, R.A., Karplus, P.A.(2018) Protein Sci 27: 573-577
- PubMed: 29139171 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3347
- Primary Citation Related Structures:
6B00 - PubMed Abstract:
Carbonic anhydrase is an enzyme of interest for many biotechnological developments including carbon sequestration. These applications often require harsh conditions, so there is a need for the development of thermostable variants. One of the most thermostable human carbonic anhydrase II (HCAIIts) variants was patented in 2006. Here, we report the ultra-high resolution crystal structure of HCAIIts. The structural changes seen are consistent with each of the six mutations involved acting largely independently and variously resulting in increased H-bonding, improved packing, and reduced side chain entropy loss on folding to yield the increased stability. We further suggest that for four of the mutations, improvements in backbone conformational energetics is also a contributor and that considerations of such conformational propensities of individual amino acids are often overlooked.
- Department of Biochemistry and Biophysics, 2011 Agriculture and Life Sciences Building, Oregon State University, Corvallis, Oregon, 97331.
Organizational Affiliation: