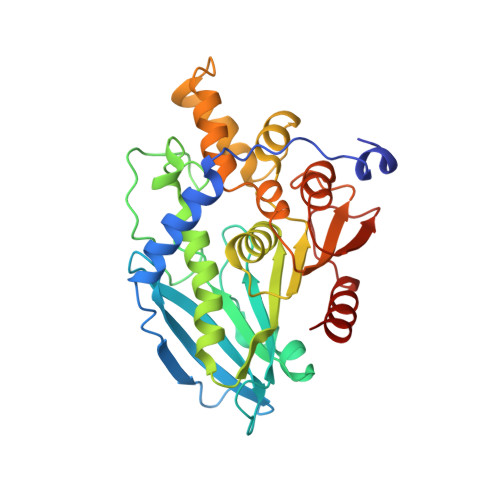
Crystal structure and functional characterization of a cold-active acetyl xylan esterase (PbAcE) from psychrophilic soil microbe Paenibacillus sp.
Park, S.H., Yoo, W., Lee, C.W., Jeong, C.S., Shin, S.C., Kim, H.W., Park, H., Kim, K.K., Kim, T.D., Lee, J.H.(2018) PLoS One 13: e0206260-e0206260
- PubMed: 30379876
- DOI: https://doi.org/10.1371/journal.pone.0206260
- Primary Citation Related Structures:
6AGQ - PubMed Abstract:
Cold-active acetyl xylan esterases allow for reduced bioreactor heating costs in bioenergy production. Here, we isolated and characterized a cold-active acetyl xylan esterase (PbAcE) from the psychrophilic soil microbe Paenibacillus sp. R4. The enzyme hydrolyzes glucose penta-acetate and xylan acetate, reversibly producing acetyl xylan from xylan, and it shows higher activity at 4°C than at 25°C. We solved the crystal structure of PbAcE at 2.1-Å resolution to investigate its active site and the reason for its low-temperature activity. Structural analysis showed that PbAcE forms a hexamer with a central substrate binding tunnel, and the inter-subunit interactions are relatively weak compared with those of its mesophilic and thermophilic homologs. PbAcE also has a shorter loop and different residue composition in the β4-α3 and β5-α4 regions near the substrate binding site. Flexible subunit movements and different active site loop conformations may enable the strong low-temperature activity and broad substrate specificity of PbAcE. In addition, PbAcE was found to have strong activity against antibiotic compound substrates, such as cefotaxime and 7-amino cephalosporanic acid (7-ACA). In conclusion, the PbAcE structure and our biochemical results provide the first example of a cold-active acetyl xylan esterase and a starting template for structure-based protein engineering.
- Unit of Polar Genomics, Korea Polar Research Institute, Incheon, Republic of Korea.
Organizational Affiliation: