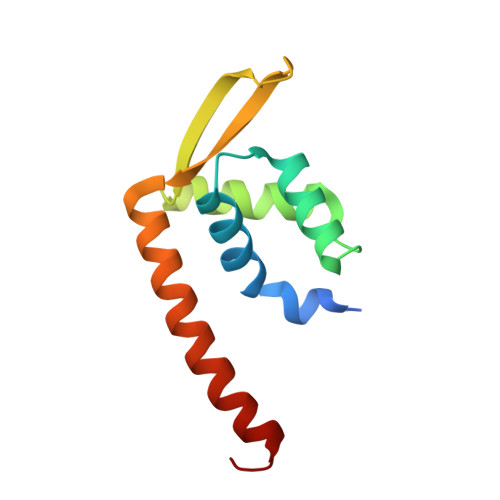
Structure-based molecular characterization and regulatory mechanism of the LftR transcription factor from Listeria monocytogenes: Conformational flexibilities and a ligand-induced regulatory mechanism.
Lee, C., Kim, M.I., Park, J., Hong, M.(2019) PLoS One 14: e0215017-e0215017
- PubMed: 30970033
- DOI: https://doi.org/10.1371/journal.pone.0215017
- Primary Citation Related Structures:
6ABQ, 6ABT - PubMed Abstract:
Listeria monocytogenes is a foodborne pathogen that causes listeriosis and can lead to serious clinical problems, such as sepsis and meningitis, in immunocompromised patients and neonates. Due to a growing number of antibiotic-resistant L. monocytogenes strains, listeriosis can steadily become refractory to antibiotic treatment. To develop novel therapeutics against listeriosis, the drug resistance mechanism of L. monocytogenes needs to be determined. The transcription factor LftR from L. monocytogenes regulates the expression of a putative multidrug resistance transporter, LieAB, and belongs to the PadR-2 subfamily of the PadR family. Despite the functional significance of LftR, our molecular understanding of the transcriptional regulatory mechanism for LftR and even for the PadR-2 subfamily is highly limited. Here, we report the crystal structure of LftR, which forms a dimer and protrudes two winged helix-turn-helix motifs for DNA recognition. Structure-based mutational and comparative analyses showed that LftR interacts with operator DNA through a LftR-specific mode as well as a common mechanism used by the PadR family. Moreover, the LftR dimer harbors one intersubunit cavity in the center of the dimeric structure as a putative ligand-binding site. Finally, conformational flexibilities in the LftR dimer and in the cavity suggest that a ligand-induced regulatory mechanism would be used by the LftR transcription factor.
- Division of Biological Science and Technology, Yonsei University, Wonju, Republic of Korea.
Organizational Affiliation: