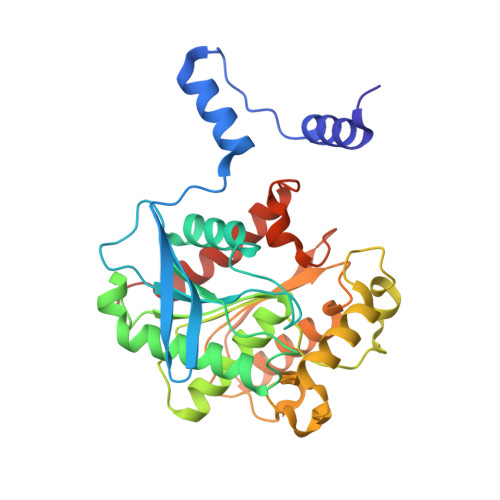
Crystal structure of chloramphenicol-metabolizing enzyme EstDL136 from a metagenome.
Kim, S.H., Kang, P.A., Han, K.T., Lee, S.W., Rhee, S.K.(2019) PLoS One 14: e0210298-e0210298
- PubMed: 30645605 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0210298
- Primary Citation Related Structures:
6AAE, 6IEY - PubMed Abstract:
Metagenomes often convey novel biological activities and therefore have gained considerable attention for use in biotechnological applications. Recently, metagenome-derived EstDL136 was found to possess chloramphenicol (Cm)-metabolizing features. Sequence analysis showed EstDL136 to be a member of the hormone-sensitive lipase (HSL) family with an Asp-His-Ser catalytic triad and a notable substrate specificity. In this study, we determined the crystal structures of EstDL136 and in a complex with Cm. Consistent with the high sequence similarity, the structure of EstDL136 is homologous to that of the HSL family. The active site of EstDL136 is a relatively shallow pocket that could accommodate Cm as a substrate as opposed to the long acyl chain substrates typical of the HSL family. Mutational analyses further suggested that several residues in the vicinity of the active site play roles in the Cm-binding of EstDL136. These results provide structural and functional insights into a metagenome-derived EstDL136.
- Department of Agricultural Biotechnology, Seoul National University, Seoul, Korea.
Organizational Affiliation: