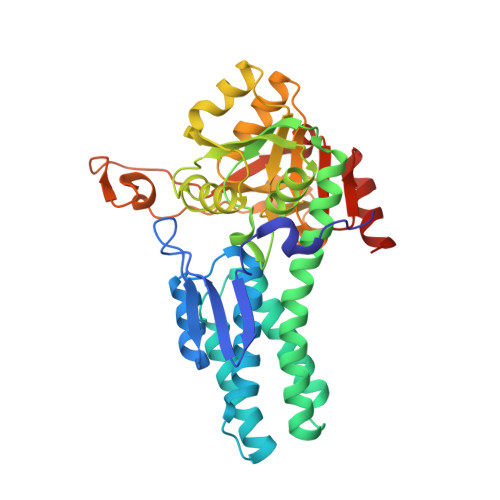
Structural insights into the catalytic mechanism of 5-methylthioribose 1-phosphate isomerase.
Gogoi, P., Mordina, P., Kanaujia, S.P.(2019) J Struct Biol 205: 67-77
- PubMed: 30471343 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2018.11.007
- Primary Citation Related Structures:
6A34, 6A35 - PubMed Abstract:
5-Methylthioribose 1-phosphate isomerase (M1Pi) is a crucial enzyme involved in the universally conserved methionine salvage pathway (MSP) where it is known to catalyze the conversion of 5-methylthioribose 1-phosphate (MTR-1-P) to 5-methylthioribulose 1-phosphate (MTRu-1-P) via a mechanism which remains unspecified till date. Furthermore, although M1Pi has a discrete function, it surprisingly shares high structural similarity with two functionally non-related proteins such as ribose-1,5-bisphosphate isomerase (R15Pi) and the regulatory subunits of eukaryotic translation initiation factor 2B (eIF2B). To identify the distinct structural features that lead to divergent functional obligations of M1Pi as well as to understand the mechanism of enzyme catalysis, the crystal structure of M1Pi from a hyperthermophilic archaeon Pyrococcus horikoshii OT3 was determined. A meticulous structural investigation of the dimeric M1Pi revealed the presence of an N-terminal extension and a hydrophobic patch absent in R15Pi and the regulatory α-subunit of eIF2B. Furthermore, unlike R15Pi in which a kink formation is observed in one of the helices, the domain movement of M1Pi is distinguished by a forward shift in a loop covering the active-site pocket. All these structural attributes contribute towards a hydrophobic microenvironment in the vicinity of the active site of the enzyme making it favorable for the reaction mechanism to commence. Thus, a hydrophobic active-site microenvironment in addition to the availability of optimal amino-acid residues surrounding the catalytic residues in M1Pi led us to propose its probable reaction mechanism via a cis-phosphoenolate intermediate formation.
- Department of Biosciences and Bioengineering, Indian Institute of Technology Guwahati, Guwahati 781039, Assam, India.
Organizational Affiliation: