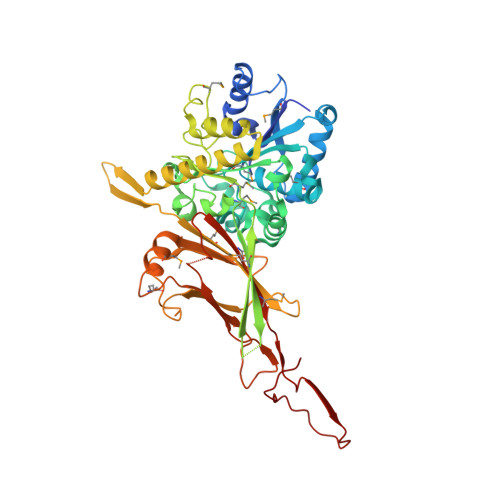
NAD(H)-mediated tetramerization controls the activity of Legionella pneumophila phospholipase PlaB.
Diwo, M., Michel, W., Aurass, P., Kuhle-Keindorf, K., Pippel, J., Krausze, J., Wamp, S., Lang, C., Blankenfeldt, W., Flieger, A.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 34074754
- DOI: https://doi.org/10.1073/pnas.2017046118
- Primary Citation Related Structures:
6ZTH, 6ZTI - PubMed Abstract:
The virulence factor PlaB promotes lung colonization, tissue destruction, and intracellular replication of Legionella pneumophila , the causative agent of Legionnaires' disease. It is a highly active phospholipase exposed at the bacterial surface and shows an extraordinary activation mechanism by tetramer deoligomerization. To unravel the molecular basis for enzyme activation and localization, we determined the crystal structure of PlaB in its tetrameric form. We found that the tetramer is a dimer of identical dimers, and a monomer consists of an N-terminal α/β-hydrolase domain expanded by two noncanonical two-stranded β-sheets, β-6/β-7 and β-9/β-10. The C-terminal domain reveals a fold displaying a bilobed β-sandwich with a hook structure required for dimer formation and structural complementation of the enzymatic domain in the neighboring monomer. This highlights the dimer as the active form. Δβ-9/β-10 mutants showed a decrease in the tetrameric fraction and altered activity profiles. The variant also revealed restricted binding to membranes resulting in mislocalization and bacterial lysis. Unexpectedly, we observed eight NAD(H) molecules at the dimer/dimer interface, suggesting that these molecules stabilize the tetramer and hence lead to enzyme inactivation. Indeed, addition of NAD(H) increased the fraction of the tetramer and concomitantly reduced activity. Together, these data reveal structural elements and an unprecedented NAD(H)-mediated tetramerization mechanism required for spatial and enzymatic control of a phospholipase virulence factor. The allosteric regulatory process identified here is suited to fine tune PlaB in a way that protects Legionella pneumophila from self-inflicted lysis while ensuring its activity at the pathogen-host interface.
- Structure and Function of Proteins, Helmholtz Centre for Infection Research, 38124 Braunschweig, Germany.
Organizational Affiliation: