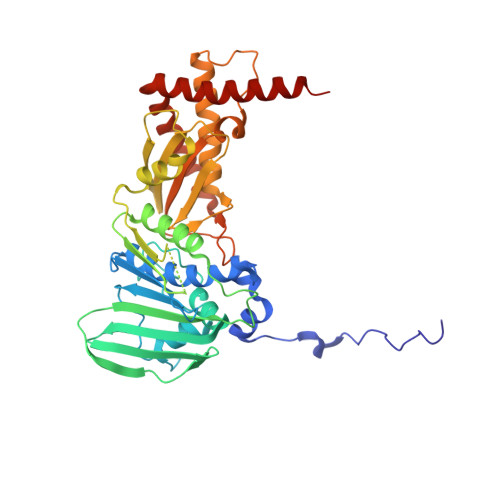
The pentapeptide-repeat protein, MfpA, interacts with mycobacterial DNA gyrase as a DNA T-segment mimic.
Feng, L., Mundy, J.E.A., Stevenson, C.E.M., Mitchenall, L.A., Lawson, D.M., Mi, K., Maxwell, A.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33836580 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2016705118
- Primary Citation Related Structures:
6ZT3, 6ZT4, 6ZT5 - PubMed Abstract:
DNA gyrase, a type II topoisomerase, introduces negative supercoils into DNA using ATP hydrolysis. The highly effective gyrase-targeted drugs, fluoroquinolones (FQs), interrupt gyrase by stabilizing a DNA-cleavage complex, a transient intermediate in the supercoiling cycle, leading to double-stranded DNA breaks. MfpA, a pentapeptide-repeat protein in mycobacteria, protects gyrase from FQs, but its molecular mechanism remains unknown. Here, we show that Mycobacterium smegmatis MfpA (MsMfpA) inhibits negative supercoiling by M. smegmatis gyrase (Msgyrase) in the absence of FQs, while in their presence, MsMfpA decreases FQ-induced DNA cleavage, protecting the enzyme from these drugs. MsMfpA stimulates the ATPase activity of Msgyrase by directly interacting with the ATPase domain (MsGyrB47), which was confirmed through X-ray crystallography of the MsMfpA-MsGyrB47 complex, and mutational analysis, demonstrating that MsMfpA mimics a T (transported) DNA segment. These data reveal the molecular mechanism whereby MfpA modulates the activity of gyrase and may provide a general molecular basis for the action of other pentapeptide-repeat proteins.
- Department of Biological Chemistry, John Innes Centre, NR4 7UH Norwich, United Kingdom.
Organizational Affiliation: