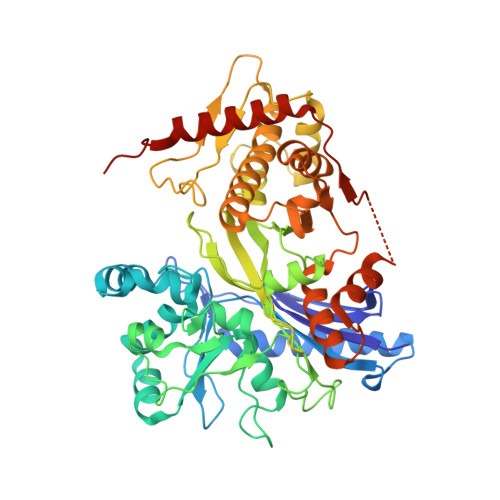
Structural Characterization of Glycerol Kinase from the Thermophilic Fungus Chaetomium thermophilum .
Wilk, P., Kuska, K., Wator, E., Malecki, P.H., Wos, K., Tokarz, P., Dubin, G., Grudnik, P.(2020) Int J Mol Sci 21
- PubMed: 33339113 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms21249570
- Primary Citation Related Structures:
6ZQ4, 6ZQ5, 6ZQ6, 6ZQ7, 6ZQ8 - PubMed Abstract:
Glycerol is an organic compound that can be utilized as an alternative source of carbon by various organisms. One of the ways to assimilate glycerol by the cell is the phosphorylative catabolic pathway in which its activation is catalyzed by glycerol kinase (GK) and glycerol-3-phosphate (G3P) is formed. To date, several GK crystal structures from bacteria, archaea, and unicellular eukaryotic parasites have been solved. Herein, we present a series of crystal structures of GK from Chaetomium thermophilum (CtGK) in apo and glycerol-bound forms. In addition, we show the feasibility of an ADP-dependent glucokinase (ADPGK)-coupled enzymatic assay to measure the CtGK activity. New structures described in our work provide structural insights into the GK catalyzed reaction in the filamentous fungus and set the foundation for understanding the glycerol metabolism in eukaryotes.
- Malopolska Centre of Biotechnology, Jagiellonian University, 7a Gronostajowa St., 30-387 Kraków, Poland.
Organizational Affiliation: