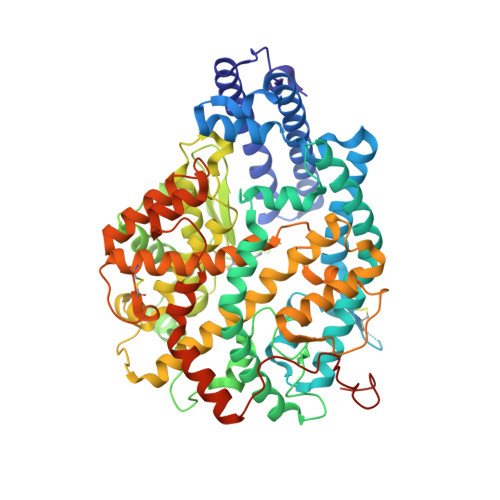
Angiotensin-converting enzyme open for business: structural insights into the subdomain dynamics.
Cozier, G.E., Lubbe, L., Sturrock, E.D., Acharya, K.R.(2021) FEBS J 288: 2238-2256
- PubMed: 33067882 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/febs.15601
- Primary Citation Related Structures:
6ZPQ, 6ZPT, 6ZPU - PubMed Abstract:
Angiotensin-1-converting enzyme (ACE) is a key enzyme in the renin-angiotensin-aldosterone and kinin systems where it cleaves angiotensin I and bradykinin peptides, respectively. However, ACE also participates in numerous other physiological functions, can hydrolyse many peptide substrates and has various exo- and endopeptidase activities. ACE achieves this complexity by containing two homologous catalytic domains (N- and C-domains), which exhibit different substrate specificities. Here, we present the first open conformation structures of ACE N-domain and a unique closed C-domain structure (2.0 Å) where the C terminus of a symmetry-related molecule is observed inserted into the active-site cavity and binding to the zinc ion. The open native N-domain structure (1.85 Å) enables comparison with ACE2, a homologue previously observed in open and closed states. An open S 2 _S'-mutant N-domain structure (2.80 Å) includes mutated residues in the S 2 and S' subsites that effect ligand binding, but are distal to the binding site. Analysis of these structures provides important insights into how structural features of the ACE domains are able to accommodate the wide variety of substrates and allow different peptidase activities. DATABASE: The atomic coordinates and structure factors for Open nACE, Open S2_S'-nACE and Native G13-cACE structures have been deposited with codes 6ZPQ, 6ZPT and 6ZPU, respectively, in the RCSB Protein Data Bank, www.pdb.org.
- Department of Biology and Biochemistry, University of Bath, Bath, UK.
Organizational Affiliation: