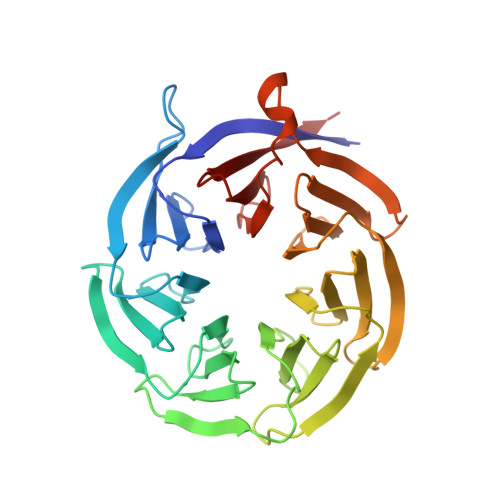
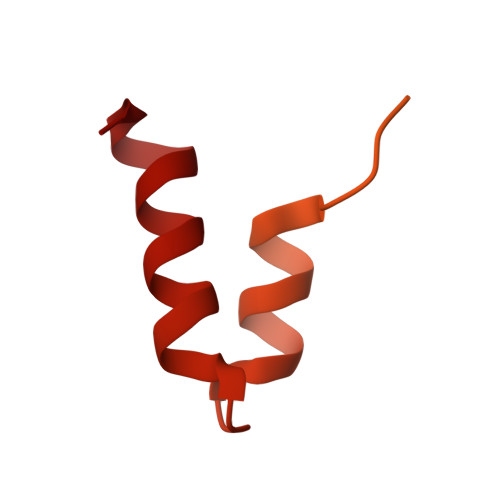
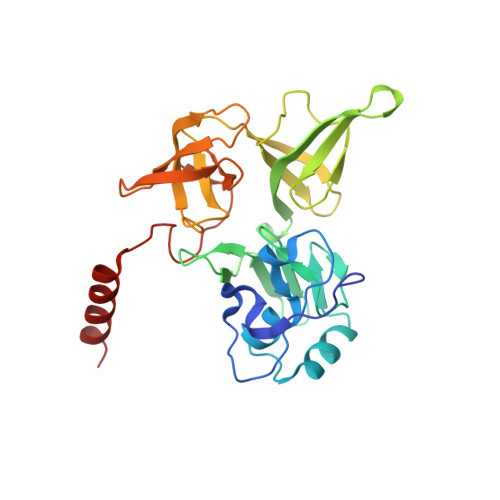
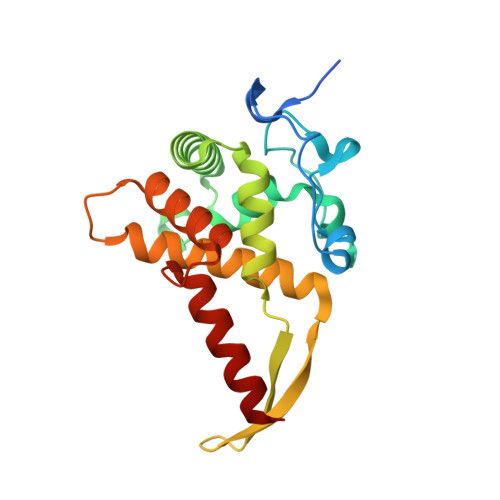
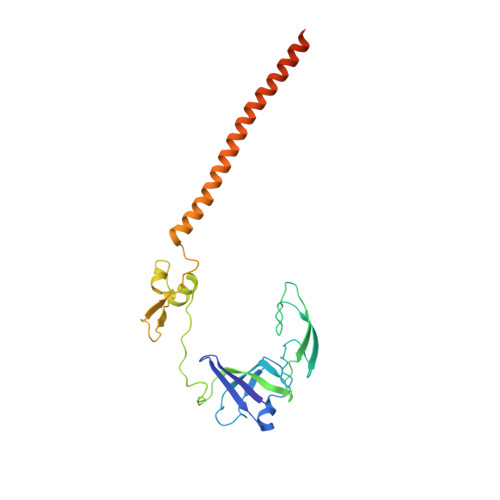
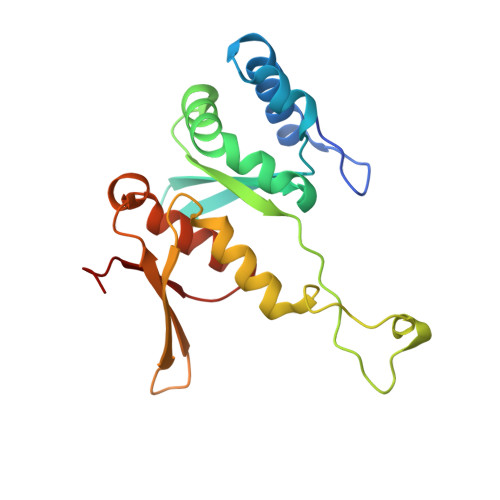
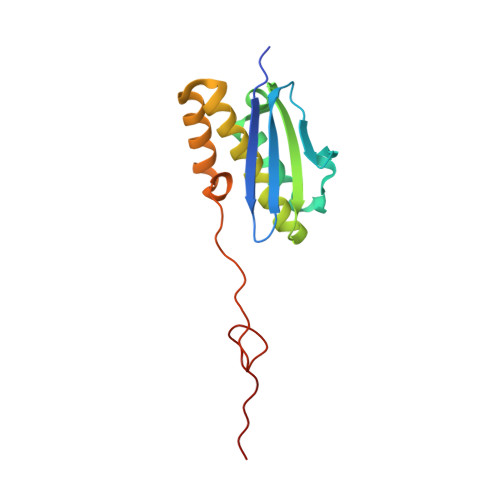
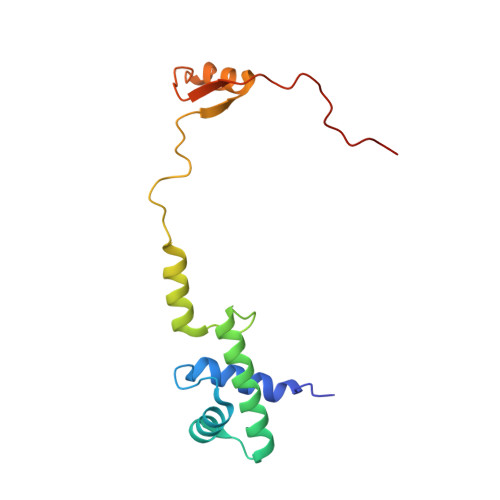
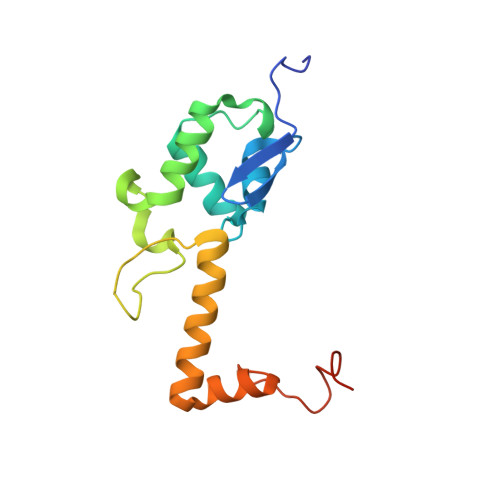
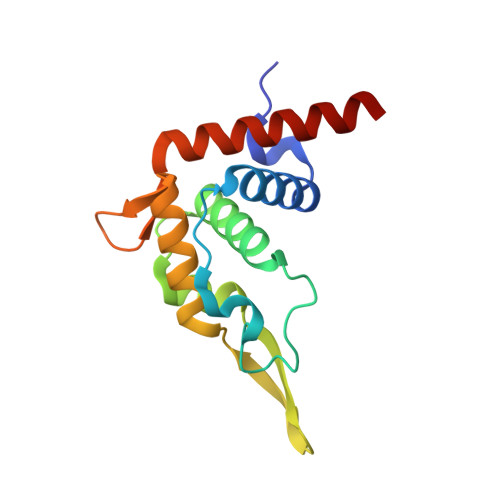
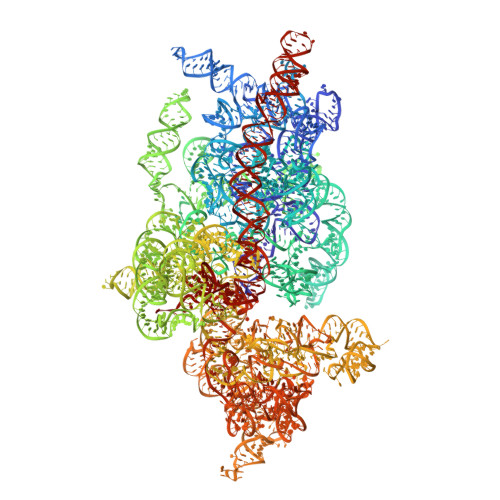
SARS-CoV-2 Nsp1 binds the ribosomal mRNA channel to inhibit translation.
Schubert, K., Karousis, E.D., Jomaa, A., Scaiola, A., Echeverria, B., Gurzeler, L.A., Leibundgut, M., Thiel, V., Muhlemann, O., Ban, N.(2020) Nat Struct Mol Biol 27: 959-966
- PubMed: 32908316 Search on PubMed
- DOI: https://doi.org/10.1038/s41594-020-0511-8
- Primary Citation Related Structures:
6ZOJ, 6ZOK, 6ZOL - PubMed Abstract:
The SARS-CoV-2 non-structural protein 1 (Nsp1), also referred to as the host shutoff factor, suppresses host innate immune functions. By combining cryo-electron microscopy and biochemistry, we show that SARS-CoV-2 Nsp1 binds to the human 40S subunit in ribosomal complexes, including the 43S pre-initiation complex and the non-translating 80S ribosome. The protein inserts its C-terminal domain into the mRNA channel, where it interferes with mRNA binding. We observe translation inhibition in the presence of Nsp1 in an in vitro translation system and in human cells. Based on the high-resolution structure of the 40S-Nsp1 complex, we identify residues of Nsp1 crucial for mediating translation inhibition. We further show that the full-length 5' untranslated region of the genomic viral mRNA stimulates translation in vitro, suggesting that SARS-CoV-2 combines global inhibition of translation by Nsp1 with efficient translation of the viral mRNA to allow expression of viral genes.
- Department of Biology, Institute of Molecular Biology and Biophysics, ETH Zurich, Zurich, Switzerland.
Organizational Affiliation: