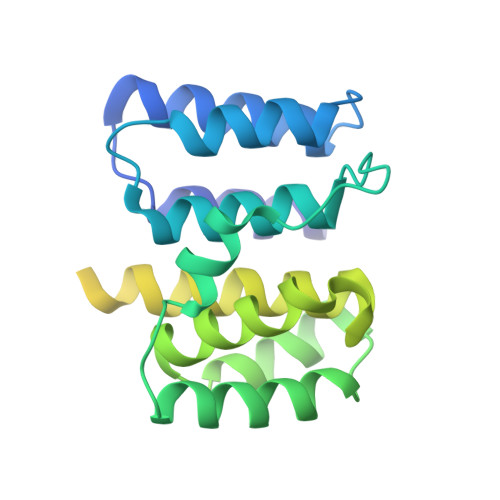
The native structure of the assembled matrix protein 1 of influenza A virus.
Peukes, J., Xiong, X., Erlendsson, S., Qu, K., Wan, W., Calder, L.J., Schraidt, O., Kummer, S., Freund, S.M.V., Krausslich, H.G., Briggs, J.A.G.(2020) Nature 587: 495-498
- PubMed: 32908308
- DOI: https://doi.org/10.1038/s41586-020-2696-8
- Primary Citation Related Structures:
6Z5J, 6Z5L - PubMed Abstract:
Influenza A virus causes millions of severe cases of disease during annual epidemics. The most abundant protein in influenza virions is matrix protein 1 (M1), which mediates virus assembly by forming an endoskeleton beneath the virus membrane 1 . The structure of full-length M1, and how it oligomerizes to mediate the assembly of virions, is unknown. Here we determine the complete structure of assembled M1 within intact virus particles, as well as the structure of M1 oligomers reconstituted in vitro. We find that the C-terminal domain of M1 is disordered in solution but can fold and bind in trans to the N-terminal domain of another M1 monomer, thus polymerizing M1 into linear strands that coat the interior surface of the membrane of the assembling virion. In the M1 polymer, five histidine residues-contributed by three different monomers of M1-form a cluster that can serve as the pH-sensitive disassembly switch after entry into a target cell. These structures therefore reveal mechanisms of influenza virus assembly and disassembly.
- Structural Studies Division, Medical Research Council Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: