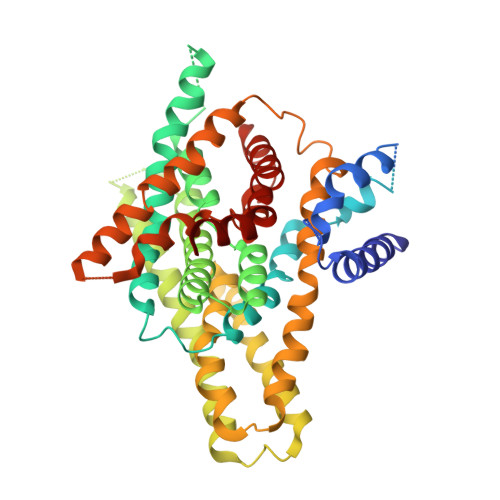
Structure and elevator mechanism of the mammalian sodium/proton exchanger NHE9.
Winklemann, I., Matsuoka, R., Meier, P.F., Shutin, D., Zhang, C., Orellana, L., Sexton, R., Landreh, M., Robinson, C.V., Beckstein, O., Drew, D.(2020) EMBO J 39: e105908-e105908
- PubMed: 33118634 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2020105908
- Primary Citation Related Structures:
6Z3Y, 6Z3Z - PubMed Abstract:
Na + /H + exchangers (NHEs) are ancient membrane-bound nanomachines that work to regulate intracellular pH, sodium levels and cell volume. NHE activities contribute to the control of the cell cycle, cell proliferation, cell migration and vesicle trafficking. NHE dysfunction has been linked to many diseases, and they are targets of pharmaceutical drugs. Despite their fundamental importance to cell homeostasis and human physiology, structural information for the mammalian NHE was lacking. Here, we report the cryogenic electron microscopy structure of NHE isoform 9 (SLC9A9) from Equus caballus at 3.2 Å resolution, an endosomal isoform highly expressed in the brain and associated with autism spectrum (ASD) and attention deficit hyperactivity (ADHD) disorders. Despite low sequence identity, the NHE9 architecture and ion-binding site are remarkably similar to distantly related bacterial Na + /H + antiporters with 13 transmembrane segments. Collectively, we reveal the conserved architecture of the NHE ion-binding site, their elevator-like structural transitions, the functional implications of autism disease mutations and the role of phosphoinositide lipids to promote homodimerization that, together, have important physiological ramifications.
- Department of Biochemistry and Biophysics, Stockholm University, Stockholm, Sweden.
Organizational Affiliation: