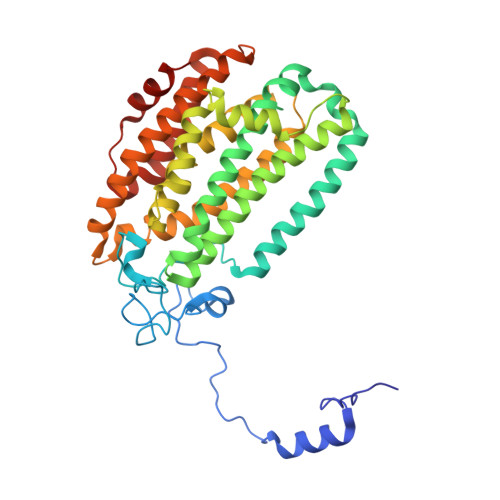
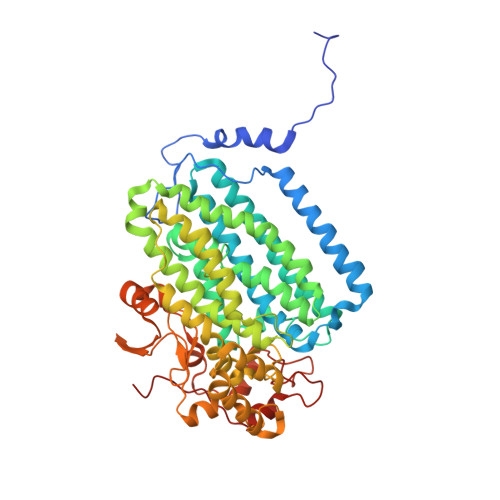
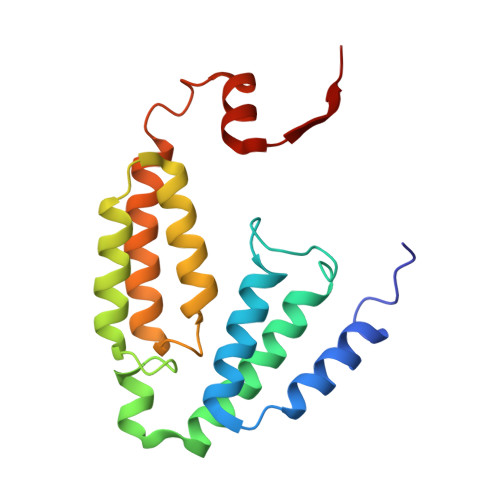
High-Resolution XFEL Structure of the Soluble Methane Monooxygenase Hydroxylase Complex with its Regulatory Component at Ambient Temperature in Two Oxidation States.
Srinivas, V., Banerjee, R., Lebrette, H., Jones, J.C., Aurelius, O., Kim, I.S., Pham, C.C., Gul, S., Sutherlin, K.D., Bhowmick, A., John, J., Bozkurt, E., Fransson, T., Aller, P., Butryn, A., Bogacz, I., Simon, P., Keable, S., Britz, A., Tono, K., Kim, K.S., Park, S.Y., Lee, S.J., Park, J., Alonso-Mori, R., Fuller, F.D., Batyuk, A., Brewster, A.S., Bergmann, U., Sauter, N.K., Orville, A.M., Yachandra, V.K., Yano, J., Lipscomb, J.D., Kern, J., Hogbom, M.(2020) J Am Chem Soc 142: 14249-14266