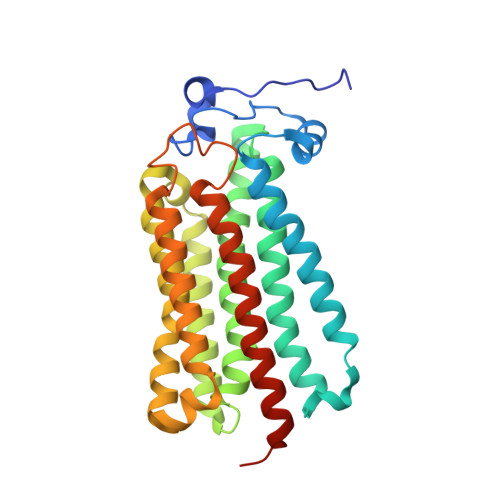
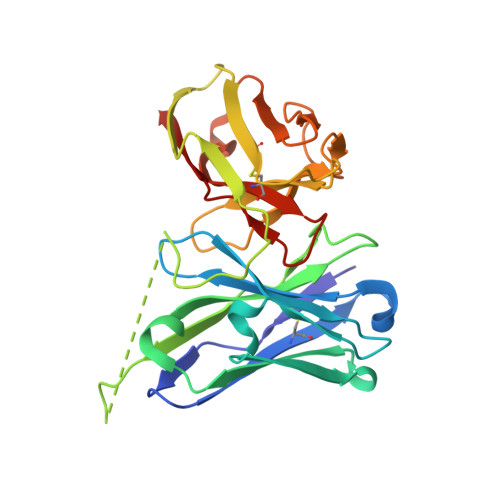
An automated platform for structural analysis of membrane proteins through serial crystallography.
Healey, R.D., Basu, S., Humm, A.S., Leyrat, C., Cong, X., Golebiowski, J., Dupeux, F., Pica, A., Granier, S., Marquez, J.A.(2021) Cell Rep Methods 1: None-None
- PubMed: 34723237 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.crmeth.2021.100102
- Primary Citation Related Structures:
6YX9, 6YXD, 6YXF, 6YXG, 6YXH - PubMed Abstract:
Membrane proteins are central to many pathophysiological processes, yet remain very difficult to analyze structurally. Moreover, high-throughput structure-based drug discovery has not yet been exploited for membrane proteins because of lack of automation. Here, we present a facile and versatile platform for in meso membrane protein crystallization, enabling rapid atomic structure determination at both cryogenic and room temperatures. We apply this approach to human integral membrane proteins, which allowed us to identify different conformational states of intramembrane enzyme-product complexes and analyze by molecular dynamics simulations the structural dynamics of the ADIPOR2 integral membrane protein. Finally, we demonstrate an automated pipeline combining high-throughput microcrystal soaking, automated laser-based harvesting, and serial crystallography, enabling screening of small-molecule libraries with membrane protein crystals grown in meso . This approach brings needed automation to this important class of drug targets and enables high-throughput structure-based ligand discovery with membrane proteins.
- IGF, University of Montpellier, CNRS, INSERM, 34094 Montpellier, France.
Organizational Affiliation: