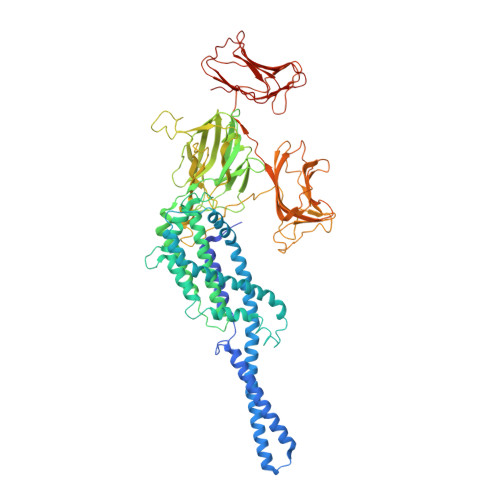
Cryo-EM structures of an insecticidal Bt toxin reveal its mechanism of action on the membrane.
Byrne, M.J., Iadanza, M.G., Perez, M.A., Maskell, D.P., George, R.M., Hesketh, E.L., Beales, P.A., Zack, M.D., Berry, C., Thompson, R.F.(2021) Nat Commun 12: 2791-2791
- PubMed: 33990582 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-23146-4
- Primary Citation Related Structures:
6YRF, 6YRG, 7NTX - PubMed Abstract:
Insect pests are a major cause of crop losses worldwide, with an estimated economic cost of $470 billion annually. Biotechnological tools have been introduced to control such insects without the need for chemical pesticides; for instance, the development of transgenic plants harbouring genes encoding insecticidal proteins. The Vip3 (vegetative insecticidal protein 3) family proteins from Bacillus thuringiensis convey toxicity to species within the Lepidoptera, and have wide potential applications in commercial agriculture. Vip3 proteins are proposed to exert their insecticidal activity through pore formation, though to date there is no mechanistic description of how this occurs on the membrane. Here we present cryo-EM structures of a Vip3 family toxin in both inactive and activated forms in conjunction with structural and functional data on toxin-membrane interactions. Together these data demonstrate that activated Vip3Bc1 complex is able to insert into membranes in a highly efficient manner, indicating that receptor binding is the likely driver of Vip3 specificity.
- Astbury Centre for Structural and Molecular Biology, School of Molecular and Cellular Biology, Faculty of Biological Sciences, University of Leeds, Leeds, UK.
Organizational Affiliation: