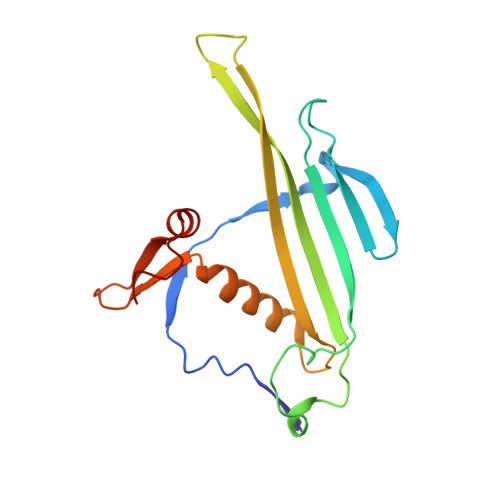
Three-dimensional structure of 22 uncultured ssRNA bacteriophages: Flexibility of the coat protein fold and variations in particle shapes.
Rumnieks, J., Lieknina, I., Kalnins, G., Sisovs, M., Akopjana, I., Bogans, J., Tars, K.(2020) Sci Adv 6
- PubMed: 32917600 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abc0023
- Primary Citation Related Structures:
6YF7, 6YF9, 6YFA, 6YFB, 6YFC, 6YFD, 6YFE, 6YFF, 6YFG, 6YFH, 6YFI, 6YFJ, 6YFK, 6YFL, 6YFM, 6YFN, 6YFO, 6YFP, 6YFQ, 6YFR, 6YFS, 6YFT, 6YFU - PubMed Abstract:
The single-stranded RNA (ssRNA) bacteriophages are among the simplest known viruses with small genomes and exceptionally high mutation rates. The number of ssRNA phage isolates has remained very low, but recent metagenomic studies have uncovered an immense variety of distinct uncultured ssRNA phages. The coat proteins (CPs) in these genomes are particularly diverse, with notable variation in length and often no recognizable similarity to previously known viruses. We recombinantly expressed metagenome-derived ssRNA phage CPs to produce virus-like particles and determined the three-dimensional structure of 22 previously uncharacterized ssRNA phage capsids covering nine distinct CP types. The structures revealed substantial deviations from the previously known ssRNA phage CP fold, uncovered an unusual prolate particle shape, and revealed a previously unseen dsRNA binding mode. These data expand our knowledge of the evolution of viral structural proteins and are of relevance for applications such as ssRNA phage-based vaccine design.
- Latvian Biomedical Research and Study Center, Rātsupītes 1, LV1067, Riga, Latvia.
Organizational Affiliation: