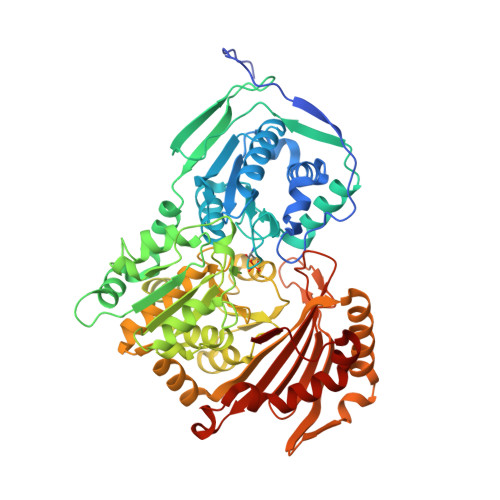
Structure and Characterization of Phosphoglucomutase 5 from Atlantic and Baltic Herring-An Inactive Enzyme with Intact Substrate Binding.
Gustafsson, R., Eckhard, U., Ye, W., Enbody, E.D., Pettersson, M., Jemth, P., Andersson, L., Selmer, M.(2020) Biomolecules 10
- PubMed: 33287293 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/biom10121631
- Primary Citation Related Structures:
6Y8X, 6Y8Y, 6Y8Z - PubMed Abstract:
Phosphoglucomutase 5 (PGM5) in humans is known as a structural muscle protein without enzymatic activity, but detailed understanding of its function is lacking. PGM5 belongs to the alpha-D-phosphohexomutase family and is closely related to the enzymatically active metabolic enzyme PGM1. In the Atlantic herring, Clupea harengus , PGM5 is one of the genes strongly associated with ecological adaptation to the brackish Baltic Sea. We here present the first crystal structures of PGM5, from the Atlantic and Baltic herring, differing by a single substitution Ala330Val. The structure of PGM5 is overall highly similar to structures of PGM1. The structure of the Baltic herring PGM5 in complex with the substrate glucose-1-phosphate shows conserved substrate binding and active site compared to human PGM1, but both PGM5 variants lack phosphoglucomutase activity under the tested conditions. Structure comparison and sequence analysis of PGM5 and PGM1 from fish and mammals suggest that the lacking enzymatic activity of PGM5 is related to differences in active-site loops that are important for flipping of the reaction intermediate. The Ala330Val substitution does not alter structure or biophysical properties of PGM5 but, due to its surface-exposed location, could affect interactions with protein-binding partners.
- Department of Cell and Molecular Biology, Uppsala University, BMC, Box 596, SE-751 24 Uppsala, Sweden.
Organizational Affiliation: