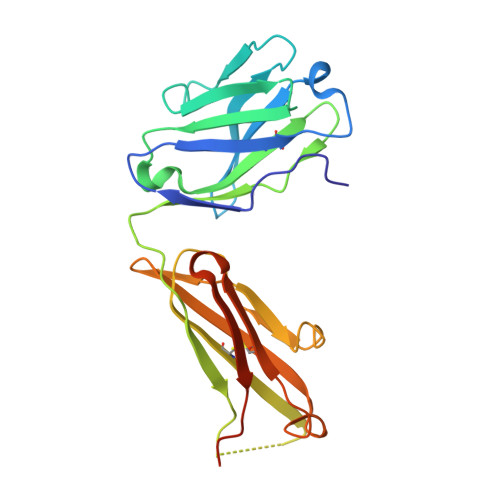
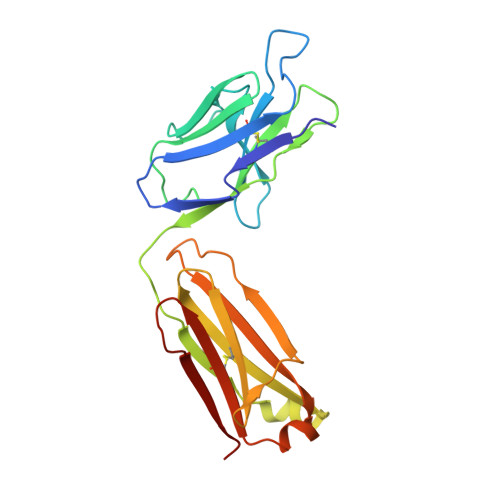
Structure of a protective epitope reveals the importance of acetylation of Neisseria meningitidis serogroup A capsular polysaccharide.
Henriques, P., Dello Iacono, L., Gimeno, A., Biolchi, A., Romano, M.R., Arda, A., Bernardes, G.J.L., Jimenez-Barbero, J., Berti, F., Rappuoli, R., Adamo, R.(2020) Proc Natl Acad Sci U S A 117: 29795-29802
- PubMed: 33158970 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2011385117
- Primary Citation Related Structures:
6Y54 - PubMed Abstract:
Meningococcal meningitis remains a substantial cause of mortality and morbidity worldwide. Until recently, countries in the African meningitis belt were susceptible to devastating outbreaks, largely attributed to serogroup A Neisseria meningitidis (MenA). Vaccination with glycoconjugates of MenA capsular polysaccharide led to an almost complete elimination of MenA clinical cases. To understand the molecular basis of vaccine-induced protection, we generated a panel of oligosaccharide fragments of different lengths and tested them with polyclonal and monoclonal antibodies by inhibition enzyme-linked immunosorbent assay, surface plasmon resonance, and competitive human serum bactericidal assay, which is a surrogate for protection. The epitope was shown to optimize between three and six repeating units and to be O -acetylated. The molecular interactions between a protective monoclonal antibody and a MenA capsular polysaccharide fragment were further elucidated at the atomic level by saturation transfer difference NMR spectroscopy and X-ray crystallography. The epitope consists of a trisaccharide anchored to the antibody via the O - and N -acetyl moieties through either H-bonding or CH-π interactions. In silico docking showed that 3- O -acetylation of the upstream residue is essential for antibody binding, while O -acetate could be equally accommodated at three and four positions of the other two residues. These results shed light on the mechanism of action of current MenA vaccines and provide a foundation for the rational design of improved therapies.
- Instituto de Medicina Molecular João Lobo Antunes, Faculdade de Medicina, Universidade de Lisboa, 1649-028 Lisboa, Portugal.
Organizational Affiliation: