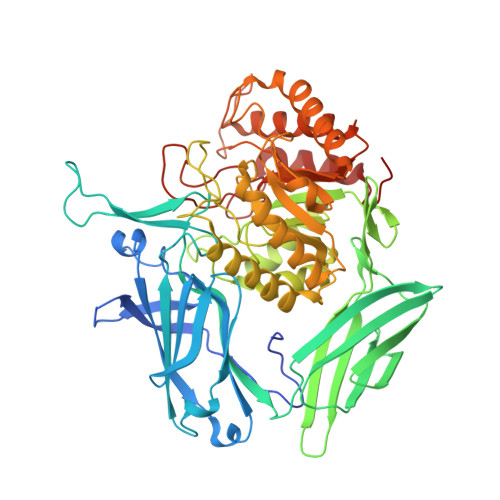
Thioglycoligation of aromatic thiols using a natural glucuronide donor.
Kurdziel, M., Kopec, M., Paris, A., Lewinski, K., Lafite, P., Daniellou, R.(2020) Org Biomol Chem 18: 5582-5585
- PubMed: 32671369 Search on PubMed
- DOI: https://doi.org/10.1039/d0ob00226g
- Primary Citation Related Structures:
6XXW - PubMed Abstract:
The β-d-glucuronidase DtGlcA from Dictyoglomus thermophilum was engineered to generate an active thioglycoligase that is able to catalyse the formation of numerous S-glucuronides. Its X-ray structure analysis indicated the ability of the biocatalyst to bind aromatic thiol acceptors for S-glycosylation. Noteworthily, the DtGlcA mutant was found to be the first thioligase that is able to use a natural sugar donor different from the widely used synthetic para-nitrophenyl glycosides.
- Institut de Chimie Organique et Analytique (ICOA), Université d'Orléans/CNRS, UMR 7311, BP 6759, F-45067, Orléans Cedex 2, France. pierre.lafite@univ-orleans.fr richard.daniellou@univ-orleans.fr.
Organizational Affiliation: