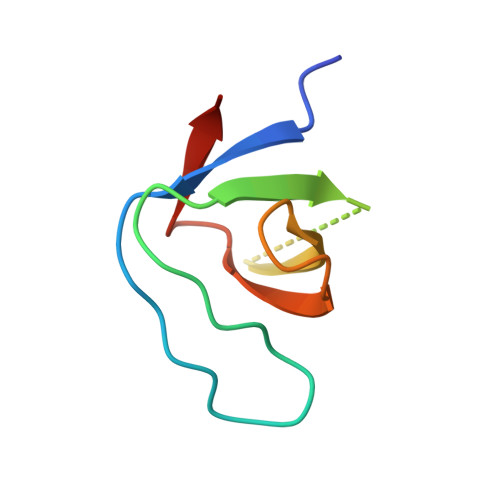
The effect of an engineered ATCUN motif on the structure and biophysical properties of the SH3 domain of c-Src tyrosine kinase.
Plaza-Garrido, M., Salinas-Garcia, M.C., Martinez, J.C., Camara-Artigas, A.(2020) J Biol Inorg Chem 25: 621-634
- PubMed: 32279137 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-020-01785-0
- Primary Citation Related Structures:
6XVM, 6XVN, 6XVO, 6XX2, 6XX3, 6XX4, 6XX5 - PubMed Abstract:
Metal binding to sites engineered in proteins can provide an increase in their stability and facilitate new functions. Besides the sites introduced in purpose, sometimes they are present accidentally as a consequence of the expression system used to produce the protein. This happens with the copper- and nickel-binding (ATCUN) motif generated by the amino-terminal residues Gly-Ser-His. This ATCUN motif is fortuitously present in many proteins, but how it affects the structural and biophysical characterization of the proteins has not been studied. In this work, we have compared the structure and biophysical properties of a small modular domain, the SH3 domain of the c-Src tyrosine kinase, cloned with and without an ATCUN motif at the N terminus. At pH 7.0, the SH3 domain with the ATCUN motif binds nickel with a binding constant K a = 28.0 ± 3.0 mM -1 . The formation of the nickel complex increases the thermal and chemical stability of the SH3 domain. A comparison of the crystal structures of the SH3 domain with and without the ATCUN motif shows that the binding of nickel does not affect the overall structure of the SH3 domain. In all crystal structures analyzed, residues Gly-Ser-His in complex with Ni 2+ show a square planar geometry. The CD visible spectrum of the nickel complex shows that this geometry is also present in the solution. Therefore, our results not only show that the ATCUN motif might influence the biophysical properties of the protein, but also points to an advantageous stabilization of the protein with potential biotechnological applications.
- Department of Chemistry and Physics, University of Almería, Agrifood Campus of International Excellence ceiA3 and CIAMBITAL, 04120, Almería, Spain.
Organizational Affiliation: