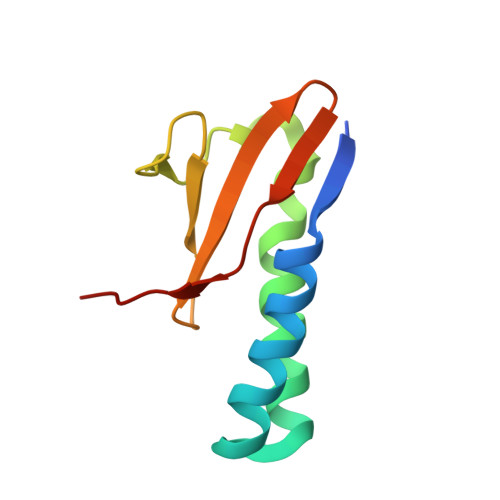
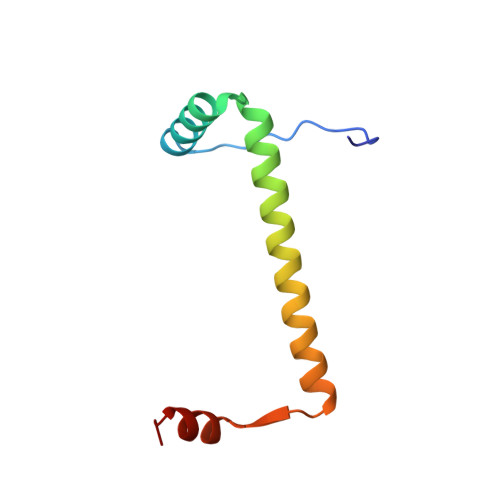
ParD Antitoxin Hotspot Alters a Disorder-to-Order Transition upon Binding to Its Cognate ParE Toxin, Lessening Its Interaction Affinity and Increasing Its Protease Degradation Kinetics.
Snead, K.J., Moore, L.L., Bourne, C.R.(2022) Biochemistry 61: 34-45
- PubMed: 34914378 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.1c00584
- Primary Citation Related Structures:
6XRW - PubMed Abstract:
Type-II toxin-antitoxin (TA) systems are comprised of two tightly interacting proteins, and operons encoding these systems have been identified throughout the genomes of bacteria. In contrast to secretion system effector-immunity pairs, TA systems must remain paired to protect the host cell from toxicity. Continual depletion of the antitoxin results in a shorter half-life than that of the toxin, though it is unclear if antitoxins can be effectively degraded when complexed with toxins. The current work probed the protein-protein interface of the PaParDE1 TA system, guided by an X-ray crystal structure, to determine contributions of antitoxin amino acids to interaction kinetics and affinity. These studies identified a "hotspot" position that alters the binding mode and resulting affinity ( K D ) from 152 pM for a 1:1 model for wild type to 25.5 and 626 nM for a 2:1 model with mutated antitoxin. This correlates with an altered induced secondary structure upon complexation with PaParE1 and increased kinetics of Lon protease digestion of the antitoxin despite the toxin presence. However, the decreased affinity at this hotspot was essentially reversed when the antitoxin dimerization region was deleted, yielding insights into complex interactions involved in the tight association. Removal of the antitoxin C-terminal seven amino acids, corresponding to the site of a disorder-to-order transition, completely prevents association. These studies combine to provide a model for the initiation of the TA interaction and highlight how manipulation of the sequence can impact the antitoxin disorder-to-order transition, weakening the affinity and resulting in increased antitoxin susceptibility to degradation.
- Department of Chemistry and Biochemistry, University of Oklahoma, 101 Stephenson Parkway, Norman, Oklahoma 73019, United States.
Organizational Affiliation: