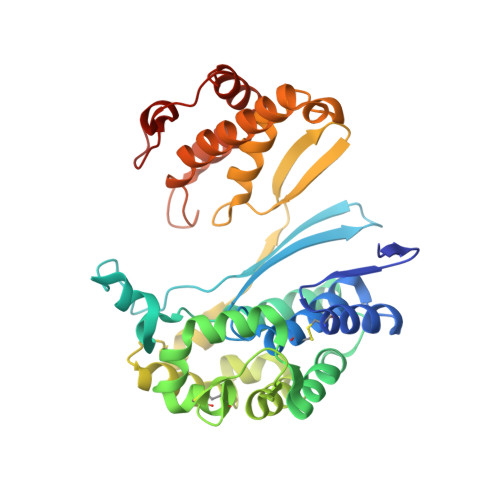
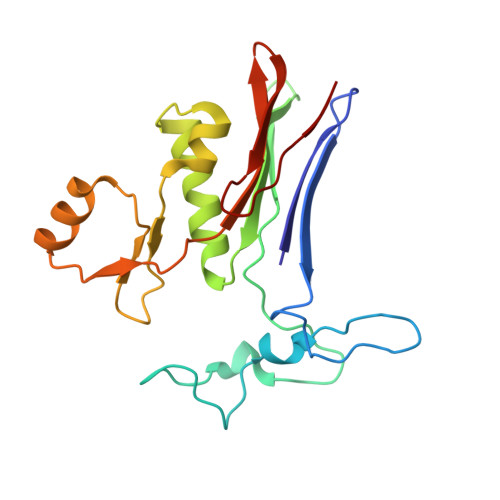
Crystal structures of glutathione- and inhibitor-bound human GGT1: critical interactions within the cysteinylglycine binding site.
Terzyan, S.S., Nguyen, L.T., Burgett, A.W.G., Heroux, A., Smith, C.A., You, Y., Hanigan, M.H.(2020) J Biological Chem 296: 100066-100066
- PubMed: 33187988 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA120.016265
- Primary Citation Related Structures:
6XPB, 6XPC - PubMed Abstract:
Overexpression of γ-glutamyl transpeptidase (GGT1) has been implicated in an array of human diseases including asthma, reperfusion injury, and cancer. Inhibitors are needed for therapy, but development of potent, specific inhibitors of GGT1 has been hampered by a lack of structural information regarding substrate binding and cleavage. To enhance our understanding of the molecular mechanism of substrate cleavage, we have solved the crystal structures of human GGT1 (hGGT1) with glutathione (a substrate) and a phosphate-glutathione analog (an irreversible inhibitor) bound in the active site. These are the first structures of any eukaryotic GGT with the cysteinylglycine region of the substrate-binding site occupied. These structures and the structure of apo-hGGT reveal movement of amino acid residues within the active site as the substrate binds. Asn-401 and Thr-381 each form hydrogen bonds with two atoms of GSH spanning the γ-glutamyl bond. Three different atoms of hGGT1 interact with the carboxyl oxygen of the cysteine of GSH. Interactions between the enzyme and substrate change as the substrate moves deeper into the active site cleft. The substrate reorients and a new hydrogen bond is formed between the substrate and the oxyanion hole. Thr-381 is locked into a single conformation as an acyl bond forms between the substrate and the enzyme. These data provide insight on a molecular level into the substrate specificity of hGGT1 and provide an explanation for seemingly disparate observations regarding the enzymatic activity of hGGT1 mutants. This knowledge will aid in the design of clinically useful hGGT1 inhibitors.
- Laboratory of Biomolecular Structure and Function, Department of Biochemistry and Molecular Biology, University of Oklahoma Health Sciences Center, Oklahoma City, Oklahoma, USA.
Organizational Affiliation: