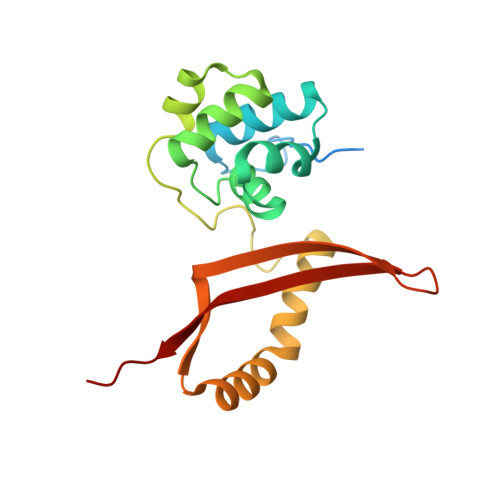
Crystal structure of a thermophilic fungal cyanase and its implications on the catalytic mechanism for bioremediation.
Ranjan, B., Choi, P.H., Pillai, S., Permaul, K., Tong, L., Singh, S.(2021) Sci Rep 11: 277-277
- PubMed: 33431973 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-020-79489-3
- Primary Citation Related Structures:
6XGT - PubMed Abstract:
Cyanase catalyzes the bicarbonate-dependent degradation of cyanate to produce ammonia and carbon dioxide, and ammonia is a considerable alternative nitrogen source. Strikingly, the cyanase from the thermophilic fungus Thermomyces lanuginosus (Tl-Cyn) has the highest catalytic efficiency reported among these enzymes. However, its molecular mechanism of action is not clearly understood, because currently there is no structural information available on fungal cyanases. Here we report the crystal structure of Tl-Cyn in complex with inhibitors malonate and formate at 2.2 Å resolution. The structure reveals extensive interactions at the subunit interfaces in a dimer, and a decamer is formed by a pentamer of these dimers. Our biochemical, kinetic and mutagenesis studies confirm the structural observations on the complex and provide further insights into its catalytic mechanism and inhibition. The structure has also aided the creation of a mutant enzyme with enhanced catalytic activity, and such enzymes may have the potential for biotechnological applications, including biotransformation and bioremediation. Moreover, other fungal cyanases with potentially high catalytic activity could also be predicted based on the Tl-Cyn structure, as the active site region among fungal cyanases are highly conserved.
- Department of Biotechnology and Food Science, Faculty of Applied Sciences, Durban University of Technology, Durban, 4000, South Africa.
Organizational Affiliation: