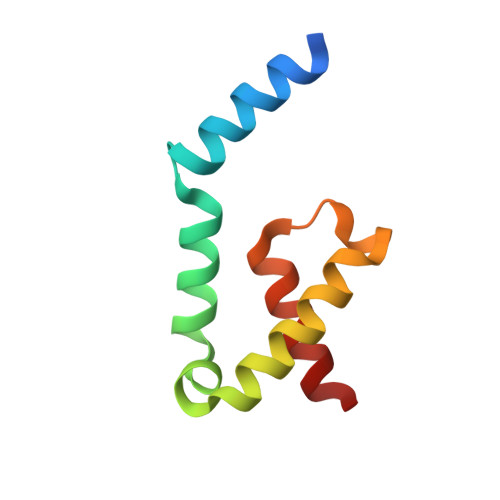
Characterization of the Pilotin-Secretin Complex from the Salmonella enterica Type III Secretion System Using Hybrid Structural Methods.
Majewski, D.D., Okon, M., Heinkel, F., Robb, C.S., Vuckovic, M., McIntosh, L.P., Strynadka, N.C.J.(2021) Structure 29: 125-138.e5
- PubMed: 32877645 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2020.08.006
- Primary Citation Related Structures:
6XFJ, 6XFK, 6XFL - PubMed Abstract:
The type III secretion system (T3SS) is a multi-membrane-spanning protein channel used by Gram-negative pathogenic bacteria to secrete effectors directly into the host cell cytoplasm. In the many species reliant on the T3SS for pathogenicity, proper assembly of the outer membrane secretin pore depends on a diverse family of lipoproteins called pilotins. We present structural and biochemical data on the Salmonella enterica pilotin InvH and the S domain of its cognate secretin InvG. Characterization of InvH by X-ray crystallography revealed a dimerized, α-helical pilotin. Size-exclusion-coupled multi-angle light scattering and small-angle X-ray scattering provide supporting evidence for the formation of an InvH homodimer in solution. Structures of the InvH-InvG heterodimeric complex determined by X-ray crystallography and NMR spectroscopy indicate a predominantly hydrophobic interface. Knowledge of the interaction between InvH and InvG brings us closer to understanding the mechanisms by which pilotins assemble the secretin pore.
- Department of Biochemistry and Molecular Biology and the Center for Blood Research, University of British Columbia, Vancouver, BC, Canada.
Organizational Affiliation: