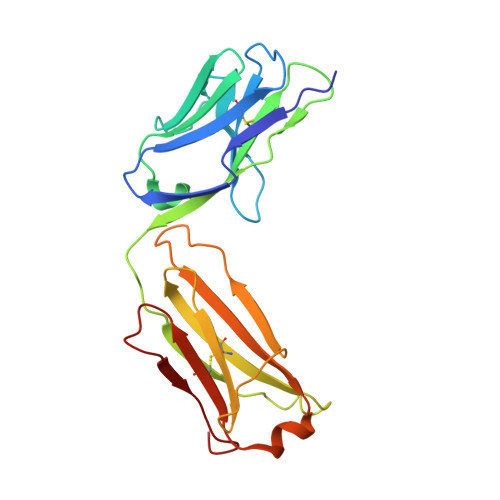
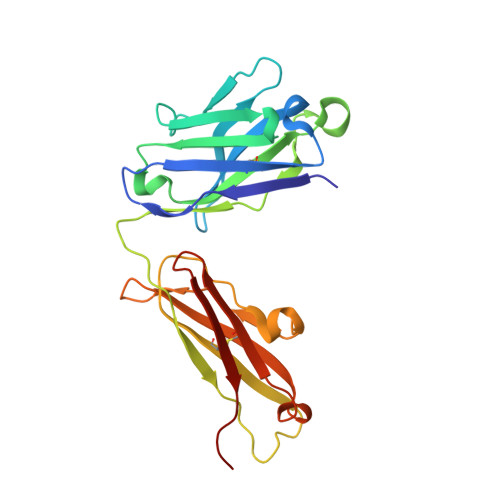
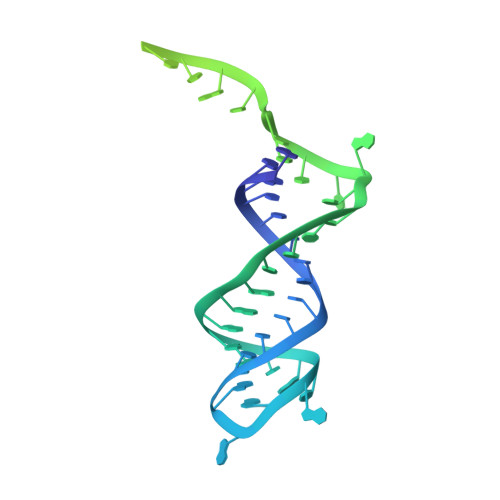
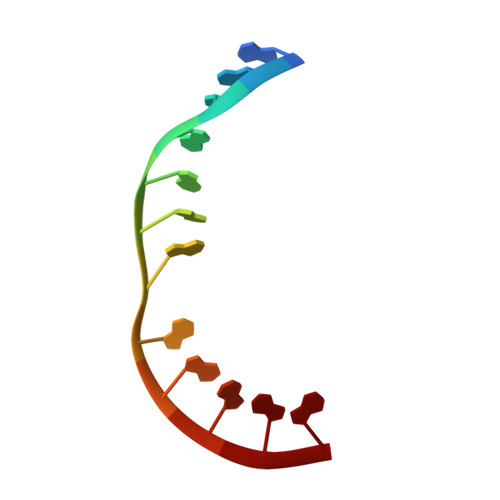
Dynamic bulge nucleotides in the KSHV PAN ENE triple helix provide a unique binding platform for small molecule ligands.
Swain, M., Ageeli, A.A., Kasprzak, W.K., Li, M., Miller, J.T., Sztuba-Solinska, J., Schneekloth, J.S., Koirala, D., Piccirili, J., Fraboni, A.J., Murelli, R.P., Wlodawer, A., Shapiro, B.A., Baird, N., Le Grice, S.F.J.(2021) Nucleic Acids Res 49: 13179-13193
- PubMed: 34871450 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab1170
- Primary Citation Related Structures:
6X5M, 6X5N - PubMed Abstract:
Cellular and virus-coded long non-coding (lnc) RNAs support multiple roles related to biological and pathological processes. Several lncRNAs sequester their 3' termini to evade cellular degradation machinery, thereby supporting disease progression. An intramolecular triplex involving the lncRNA 3' terminus, the element for nuclear expression (ENE), stabilizes RNA transcripts and promotes persistent function. Therefore, such ENE triplexes, as presented here in Kaposi's sarcoma-associated herpesvirus (KSHV) polyadenylated nuclear (PAN) lncRNA, represent targets for therapeutic development. Towards identifying novel ligands targeting the PAN ENE triplex, we screened a library of immobilized small molecules and identified several triplex-binding chemotypes, the tightest of which exhibits micromolar binding affinity. Combined biophysical, biochemical, and computational strategies localized ligand binding to a platform created near a dinucleotide bulge at the base of the triplex. Crystal structures of apo (3.3 Å) and ligand-soaked (2.5 Å) ENE triplexes, which include a stabilizing basal duplex, indicate significant local structural rearrangements within this dinucleotide bulge. MD simulations and a modified nucleoside analog interference technique corroborate the role of the bulge and the base of the triplex in ligand binding. Together with recently discovered small molecules that reduce nuclear MALAT1 lncRNA levels by engaging its ENE triplex, our data supports the potential of targeting RNA triplexes with small molecules.
- Basic Research Laboratory, National Cancer Institute, Frederick, MD 21702, USA.
Organizational Affiliation: