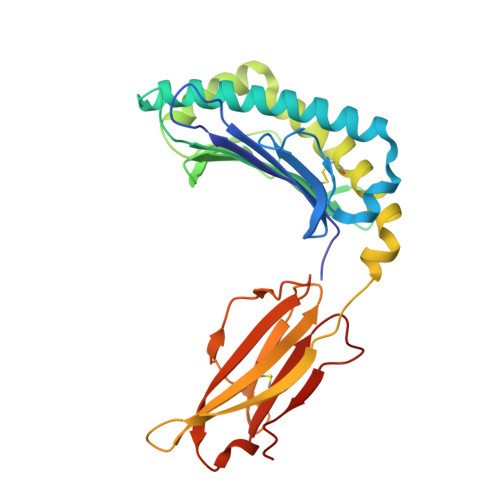
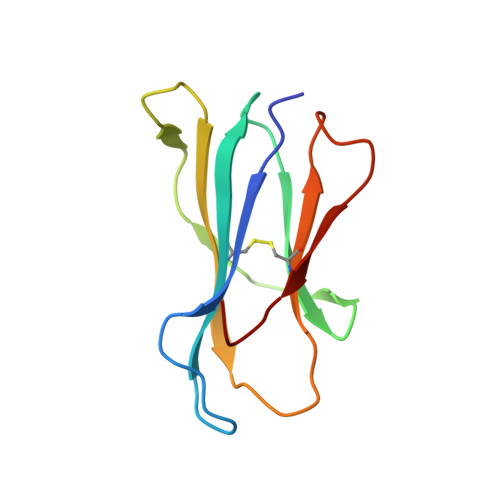
Overlapping Peptides Elicit Distinct CD8 + T Cell Responses following Influenza A Virus Infection.
Assmus, L.M., Guan, J., Wu, T., Farenc, C., Sng, X.Y.X., Zareie, P., Nguyen, A., Nguyen, A.T., Tscharke, D.C., Thomas, P.G., Rossjohn, J., Gras, S., Croft, N.P., Purcell, A.W., La Gruta, N.L.(2020) J Immunol 205: 1731-1742
- PubMed: 32868409 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4049/jimmunol.2000689
- Primary Citation Related Structures:
6WZY, 6X00 - PubMed Abstract:
The presentation of pathogen-derived peptides on MHC class I molecules is essential for the initiation of adaptive CD8 + T cell immunity, which in turn is critical for effective control of many significant human infections. The identification of immunogenic pathogen-derived epitopes and a detailed understanding of how they are recognized by TCRs is essential for the design of effective T cell-based vaccines. In this study, we have characterized the T cell recognition and immune responses in mice to two naturally presented influenza A virus-derived peptides previously identified from virally infected cells via mass spectrometry. These neuraminidase-derived peptides, NA 181-190 (SGPDNGAVAV) and NA 181-191 (SGPDNGAVAVL), are completely overlapping with the exception of a 1 aa extension at the C terminus of the longer peptide. This minor peptidic difference results in the induction of two completely independent and non-cross-reactive T cell populations that show distinct functional characteristics after influenza A virus infection of B6 mice. We show that the unique TCR reactivity to the overlapping peptides is present in the naive repertoire prior to immune expansion in B6 mice. Moreover, we provide a structural explanation underlying the distinct CD8 + T cell reactivities, which reinforces the concept that peptide length is a key determinant of Ag specificity in CD8 + T cell responses.
- Department of Biochemistry and Molecular Biology, Monash Biomedicine Discovery Institute, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: