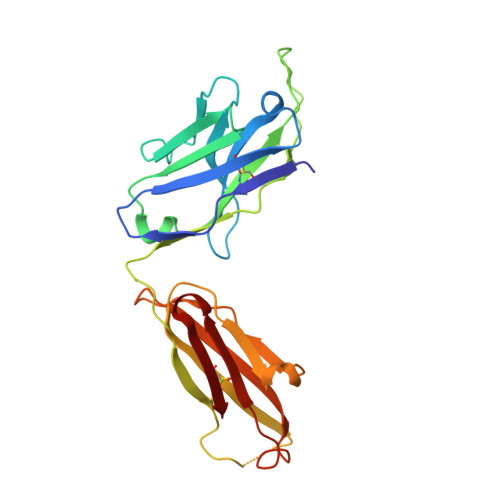
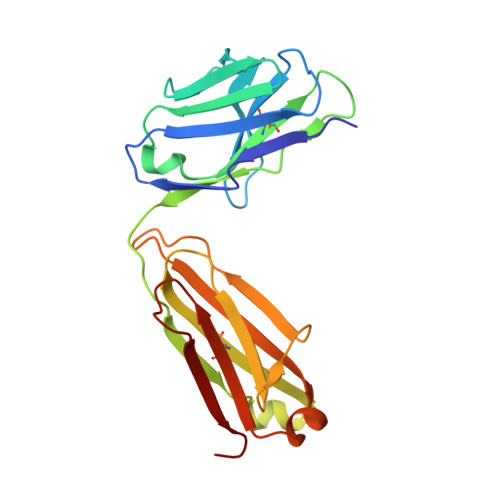
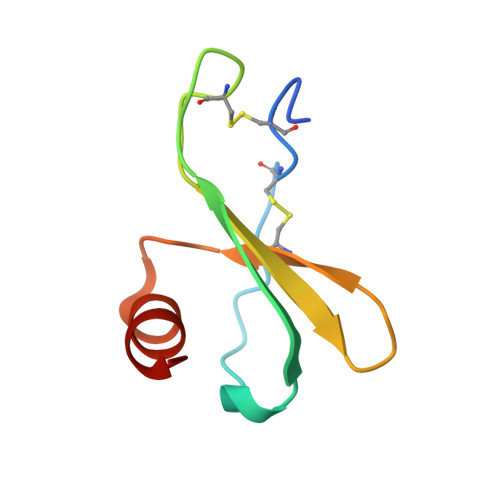
Discovery and characterization of a neutralizing pan-ELR+CXC chemokine monoclonal antibody.
Boyles, J.S., Beidler, C.B., Strifler, B.A., Girard, D.S., Druzina, Z., Durbin, J.D., Swearingen, M.L., Lee, L.N., Kikly, K., Chintharlapalli, S., Witcher, D.R.(null) MAbs 12: 1831880-1831880
- PubMed: 33183151 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/19420862.2020.1831880
- Primary Citation Related Structures:
6WZJ, 6WZK, 6WZL, 6WZM - PubMed Abstract:
CXCR1 and CXCR2 signaling play a critical role in neutrophil migration, angiogenesis, and tumorigenesis and are therefore an attractive signaling axis to target in a variety of indications. In human, a total of seven chemokines signal through these receptors and comprise the ELR + CXC chemokine family, so named because of the conserved ELRCXC N-terminal motif. To fully antagonize CXCR1 and CXCR2 signaling, an effective therapeutic should block either both receptors or all seven ligands, yet neither approach has been fully realized clinically. In this work, we describe the generation and characterization of LY3041658, a humanized monoclonal antibody that binds and neutralizes all seven human and cynomolgus monkey ELR + CXC chemokines and three of five mouse and rat ELR + CXC chemokines with high affinity. LY3041658 is able to block ELR + CXC chemokine-induced Ca 2+ mobilization, CXCR2 internalization, and chemotaxis in vitro as well as neutrophil mobilization in vivo without affecting other neutrophil functions. In addition to the in vitro and in vivo activity, we characterized the epitope and structural basis for binding in detail through alanine scanning, crystallography, and mutagenesis. Together, these data provide a robust preclinical characterization of LY3041658 for which the efficacy and safety is being evaluated in human clinical trials for neutrophilic skin diseases.
- Biotechnology Discovery Research, Lilly Research Laboratories, Eli Lilly and Company , Indianapolis, IN, USA.
Organizational Affiliation: