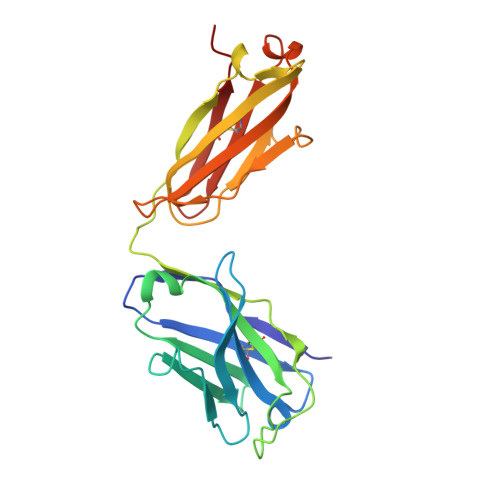
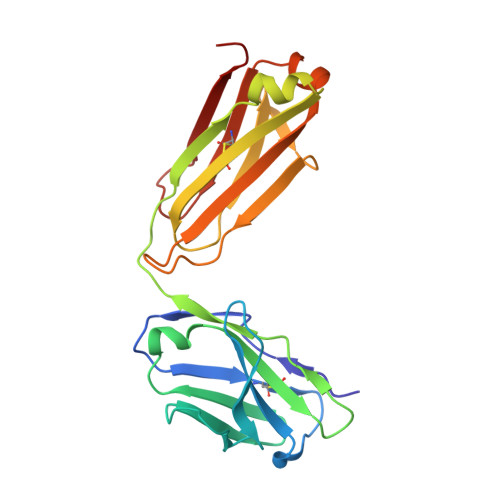
Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody.
Pinto, D., Park, Y.J., Beltramello, M., Walls, A.C., Tortorici, M.A., Bianchi, S., Jaconi, S., Culap, K., Zatta, F., De Marco, A., Peter, A., Guarino, B., Spreafico, R., Cameroni, E., Case, J.B., Chen, R.E., Havenar-Daughton, C., Snell, G., Telenti, A., Virgin, H.W., Lanzavecchia, A., Diamond, M.S., Fink, K., Veesler, D., Corti, D.(2020) Nature 583: 290-295
- PubMed: 32422645 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-020-2349-y
- Primary Citation Related Structures:
6WPS, 6WPT, 6WS6 - PubMed Abstract:
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a newly emerged coronavirus that is responsible for the current pandemic of coronavirus disease 2019 (COVID-19), which has resulted in more than 3.7 million infections and 260,000 deaths as of 6 May 2020 1,2 . Vaccine and therapeutic discovery efforts are paramount to curb the pandemic spread of this zoonotic virus. The SARS-CoV-2 spike (S) glycoprotein promotes entry into host cells and is the main target of neutralizing antibodies. Here we describe several monoclonal antibodies that target the S glycoprotein of SARS-CoV-2, which we identified from memory B cells of an individual who was infected with severe acute respiratory syndrome coronavirus (SARS-CoV) in 2003. One antibody (named S309) potently neutralizes SARS-CoV-2 and SARS-CoV pseudoviruses as well as authentic SARS-CoV-2, by engaging the receptor-binding domain of the S glycoprotein. Using cryo-electron microscopy and binding assays, we show that S309 recognizes an epitope containing a glycan that is conserved within the Sarbecovirus subgenus, without competing with receptor attachment. Antibody cocktails that include S309 in combination with other antibodies that we identified further enhanced SARS-CoV-2 neutralization, and may limit the emergence of neutralization-escape mutants. These results pave the way for using S309 and antibody cocktails containing S309 for prophylaxis in individuals at a high risk of exposure or as a post-exposure therapy to limit or treat severe disease.
- Humabs BioMed SA, Vir Biotechnology, Bellinzona, Switzerland.
Organizational Affiliation: