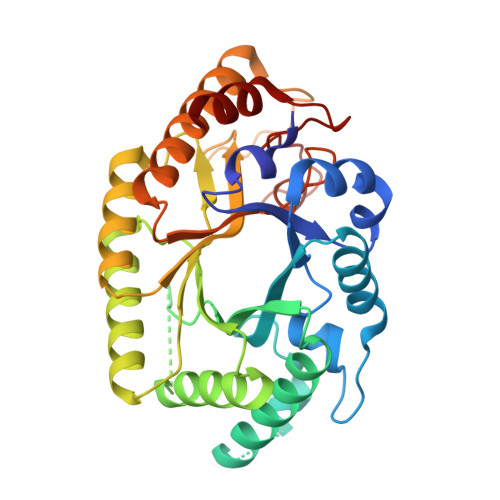
Transformation of xylan into value-added biocommodities using Thermobacillus composti GH10 xylanase.
Sepulchro, A.G.V., Pellegrini, V.O.A., Briganti, L., de Araujo, E.A., de Araujo, S.S., Polikarpov, I.(2020) Carbohydr Polym 247: 116714-116714
- PubMed: 32829841 Search on PubMed
- DOI: https://doi.org/10.1016/j.carbpol.2020.116714
- Primary Citation Related Structures:
6WQW - PubMed Abstract:
Enzymatic transformation of xylans into renewable fuels and value-added products is mediated by xylanases. Here we describe the biochemical and X-ray structural characterization of Thermobacillus composti GH10 xylanase (TcXyn10A) at 2.1 Å resolution aiming to unravel details of its recognition of glucurono- and arabinoxylan at a molecular level. TcXyn10A improves the efficiency of pretreated lignocellulosic biomass hydrolysis by a commercial enzyme cocktail causing a 15.35 % increase in xylose release and 4.38 % glucose release after 24 h of reaction. The enzyme releases predominantly xylobiose and xylotriose, as well as MeGlcA3 × 3 (from beechwood glucuronoxylan) and a range of decorated xylooligosaccharides (XOS) from rye arabinoxylan, with Ara2 × 2 being the major product. The enzyme liberates XOS with the yields of 29.09 % for beechwood glucuronoxylan and 16.98 % for rye arabinoxylan. Finally, TcXyn10A has a high thermal stability, halotolerance, and resistance to ethanol, biochemical properties that can be desirable for a number of industrial applications.
- Instituto de Física de São Carlos, Universidade de São Paulo, Avenida Trabalhador São-carlense 400, 13566-590, São Carlos, SP, Brazil.
Organizational Affiliation: