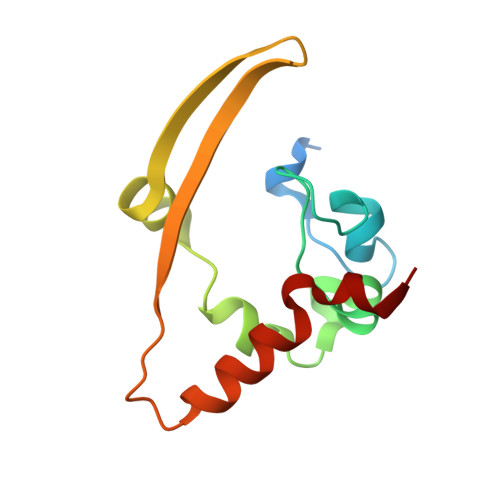
Serodominant SARS-CoV-2 Nucleocapsid Peptides Map to Unstructured Protein Regions.
Vandervaart, J.P., Inniss, N.L., Ling-Hu, T., Minasov, G., Wiersum, G., Rosas-Lemus, M., Shuvalova, L., Achenbach, C.J., Hultquist, J.F., Satchell, K.J.F., Bachta, K.E.R.(2023) Microbiol Spectr : e0032423-e0032423
- PubMed: 37191546
- DOI: https://doi.org/10.1128/spectrum.00324-23
- Primary Citation Related Structures:
6WJI - PubMed Abstract:
The SARS-CoV-2 nucleocapsid (N) protein is highly immunogenic, and anti-N antibodies are commonly used as markers for prior infection. While several studies have examined or predicted the antigenic regions of N, these have lacked consensus and structural context. Using COVID-19 patient sera to probe an overlapping peptide array, we identified six public and four private epitope regions across N, some of which are unique to this study. We further report the first deposited X-ray structure of the stable dimerization domain at 2.05 Å as similar to all other reported structures. Structural mapping revealed that most epitopes are derived from surface-exposed loops on the stable domains or from the unstructured linker regions. An antibody response to an epitope in the stable RNA binding domain was found more frequently in sera from patients requiring intensive care. Since emerging amino acid variations in N map to immunogenic peptides, N protein variation could impact detection of seroconversion for variants of concern. IMPORTANCE As SARS-CoV-2 continues to evolve, a structural and genetic understanding of key viral epitopes will be essential to the development of next-generation diagnostics and vaccines. This study uses structural biology and epitope mapping to define the antigenic regions of the viral nucleocapsid protein in sera from a cohort of COVID-19 patients with diverse clinical outcomes. These results are interpreted in the context of prior structural and epitope mapping studies as well as in the context of emergent viral variants. This report serves as a resource for synthesizing the current state of the field toward improving strategies for future diagnostic and therapeutic design.
- Department of Microbiology-Immunology, Northwestern University, Feinberg School of Medicine, Chicago, Illinois, USA.
Organizational Affiliation: