Structure of a Hallucinogen-Activated Gq-Coupled 5-HT2A Serotonin Receptor
Kim, K., Che, T., Panova, O., Shoichet, B.K., Skiniotis, G., Roth, B.L.(2020) Cell 182: 1574-1588
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
(2020) Cell 182: 1574-1588
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Soluble cytochrome b562,5-hydroxytryptamine receptor 2A chimera | 497 | Escherichia coli, Homo sapiens This entity is chimeric | Mutation(s): 0 Gene Names: cybC, HTR2A, HTR2 Membrane Entity: Yes | 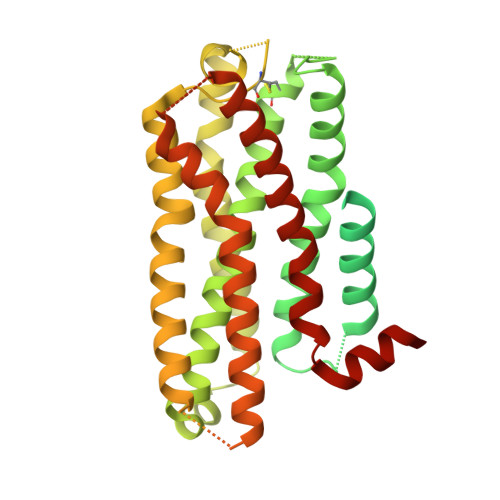 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P28223 GTEx: ENSG00000102468 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | P28223P0ABE7 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| G subunit q (Gi2-mini-Gq chimeric) | 246 | Homo sapiens | Mutation(s): 0 | 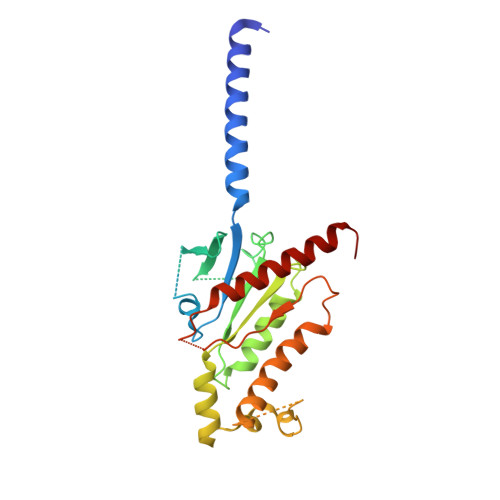 | |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 | 358 | Homo sapiens | Mutation(s): 0 Gene Names: GNB1 | 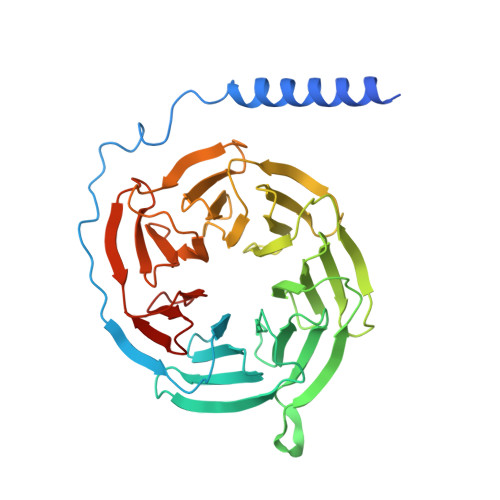 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P62873 GTEx: ENSG00000078369 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P62873 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 | 71 | Homo sapiens | Mutation(s): 0 Gene Names: GNG2 Membrane Entity: Yes | 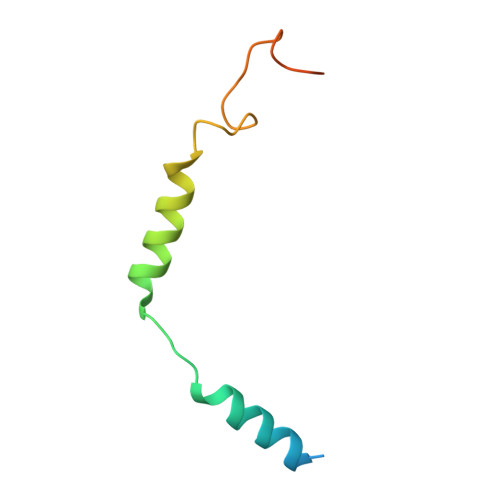 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P59768 GTEx: ENSG00000186469 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P59768 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| single Fab chain (svFv16) | 267 | Homo sapiens | Mutation(s): 0 |  | |
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| U0G (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | F [auth A] | 4-(2-{[(2-hydroxyphenyl)methyl]amino}ethyl)-2,5-dimethoxybenzonitrile C18 H20 N2 O3 VWEDZTZAXHMZIL-UHFFFAOYSA-N |  | ||
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute on Drug Abuse (NIH/NIDA) | United States | R37 DA045657 |
| National Institutes of Health/National Institute of Mental Health (NIH/NIMH) | United States | RO1 MH112205 |