The crystal structure of a collagen galactosylhydroxylysyl glucosyltransferase from human
Guo, H.-F., Tsai, C.-L., Miller, M.D., Phillips Jr., G.N., Tainer, J.A., Kurie, J.M.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Multifunctional procollagen lysine hydroxylase and glycosyltransferase LH3 | 247 | Homo sapiens | Mutation(s): 0 Gene Names: PLOD3 EC: 1.14.11.4 (PDB Primary Data), 2.4.1.50 (PDB Primary Data), 2.4.1.66 (PDB Primary Data) | 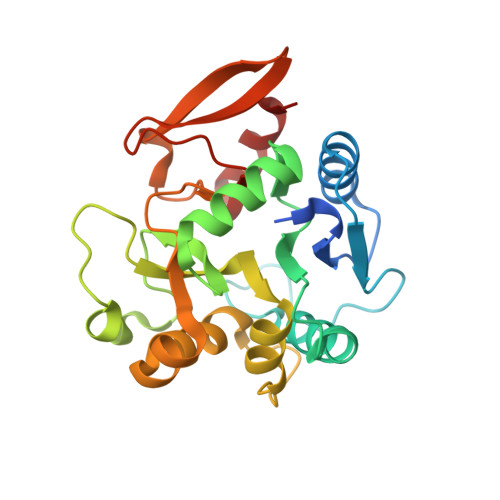 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O60568 GTEx: ENSG00000106397 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O60568 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| UDP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth A] | URIDINE-5'-DIPHOSPHATE C9 H14 N2 O12 P2 XCCTYIAWTASOJW-XVFCMESISA-N |  | ||
| TRS Download:Ideal Coordinates CCD File | F [auth A] | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL C4 H12 N O3 LENZDBCJOHFCAS-UHFFFAOYSA-O |  | ||
| MN (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | B [auth A] | MANGANESE (II) ION Mn WAEMQWOKJMHJLA-UHFFFAOYSA-N |  | ||
| MG (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A], D [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 71.066 | α = 90 |
| b = 71.066 | β = 90 |
| c = 110.813 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| SHELXD | phasing |
| SOLVE | phasing |
| RESOLVE | model building |
| SHELXE | model building |
| Coot | model building |
| PDB_EXTRACT | data extraction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Cancer Institute (NIH/NCI) | United States | R01CA105155 |
| National Institutes of Health/National Cancer Institute (NIH/NCI) | United States | K99CA225633 |
| Cancer Prevention and Research Institute of Texas (CPRIT) | United States | RP160652 |