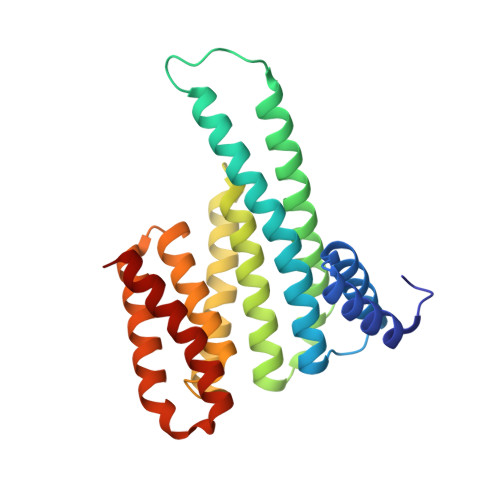
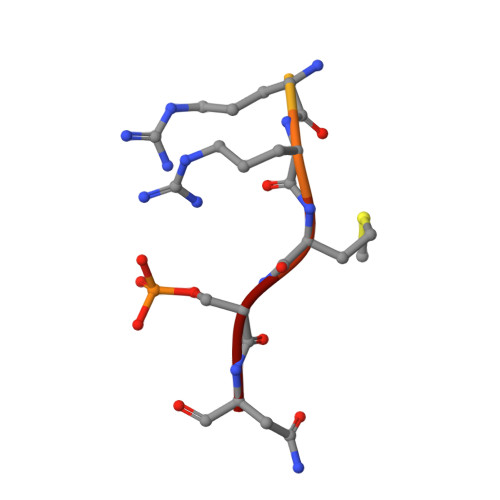
Henipavirus W Proteins Interact with 14-3-3 To Modulate Host Gene Expression.
Edwards, M.R., Hoad, M., Tsimbalyuk, S., Menicucci, A.R., Messaoudi, I., Forwood, J.K., Basler, C.F.(2020) J Virol 94
- PubMed: 32321809 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00373-20
- Primary Citation Related Structures:
6W0L - PubMed Abstract:
Nipah virus (NiV) and Hendra virus (HeV), members of the Henipavirus genus in the Paramyxoviridae family, are recently emerged, highly lethal zoonotic pathogens. The NiV and HeV nonsegmented, negative-sense RNA genomes encode nine proteins, including the W protein. Expressed from the P gene through mRNA editing, W shares a common N-terminus with P and V but has a unique C-terminus. Expressed alone, W modulates innate immune responses by several mechanisms, and elimination of W from NiV alters the course of infection in experimentally infected ferrets. However, the specific host interactions that allow W to modulate innate immunity are incompletely understood. This study demonstrates that the NiV and HeV W proteins interact with all seven isoforms of the 14-3-3 family, regulatory molecules that preferentially bind phosphorylated target proteins to regulate a wide range of cellular functions. The interaction is dependent on the penultimate amino acid residue in the W sequence, a conserved, phosphorylated serine. The cocrystal structure of the W C-terminal binding motif with 14-3-3 provides only the second structure of a complex containing a mode III interactor, which is defined as a 14-3-3 interaction with a phosphoserine/phosphothreonine at the C-termini of the target protein. Transcriptomic analysis of inducible cell lines infected with an RNA virus and expressing either wild-type W or W lacking 14-3-3 binding, identifies new functions for W. These include the regulation of cellular metabolic processes, extracellular matrix organization, and apoptosis. IMPORTANCE Nipah virus (NiV) and Hendra virus (HeV), members of the Henipavirus genus, are recently emerged, highly lethal zoonotic pathogens that cause yearly outbreaks. NiV and HeV each encode a W protein that has roles in regulating host signaling pathways, including antagonism of the innate immune response. However, the mechanisms used by W to regulate these host responses are not clear. Here, characterization of the interaction of NiV and HeV W with 14-3-3 identifies modulation of 14-3-3-regulated host signaling pathways not previously associated with W, suggesting new avenues of research. The cocrystal structure of the NiV W:14-3-3 complex, as only the second structure of a 14-3-3 mode III interactor, provides further insight into this less-well-understood 14-3-3 binding motif.
- Center for Microbial Pathogenesis, Institute for Biomedical Sciences, Georgia State University, Atlanta, Georgia, USA.
Organizational Affiliation: