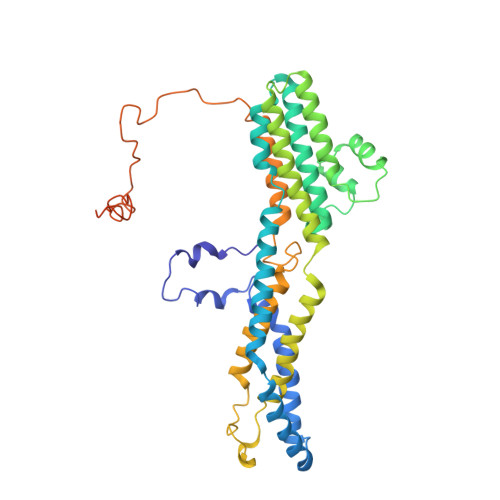
Structural and functional characterization of the bestrophin-2 anion channel.
Owji, A.P., Zhao, Q., Ji, C., Kittredge, A., Hopiavuori, A., Fu, Z., Ward, N., Clarke, O.B., Shen, Y., Zhang, Y., Hendrickson, W.A., Yang, T.(2020) Nat Struct Mol Biol 27: 382-391
- PubMed: 32251414 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-020-0402-z
- Primary Citation Related Structures:
6VX5, 6VX6, 6VX7, 6VX8, 6VX9 - PubMed Abstract:
The bestrophin family of calcium (Ca 2+ )-activated chloride (Cl - ) channels, which mediate the influx and efflux of monovalent anions in response to the levels of intracellular Ca 2+ , comprises four members in mammals (bestrophin 1-4). Here we report cryo-EM structures of bovine bestrophin-2 (bBest2) bound and unbound by Ca 2+ at 2.4- and 2.2-Å resolution, respectively. The bBest2 structure highlights four previously underappreciated pore-lining residues specifically conserved in Best2 but not in Best1, illustrating the differences between these paralogs. Structure-inspired electrophysiological analysis reveals that, although the channel is sensitive to Ca 2+ , it has substantial Ca 2+ -independent activity for Cl - , reflecting the opening at the cytoplasmic restriction of the ion conducting pathway even when Ca 2+ is absent. Moreover, the ion selectivity of bBest2 is controlled by multiple residues, including those involved in gating.
- Department of Pharmacology, Columbia University, New York, NY, USA.
Organizational Affiliation: