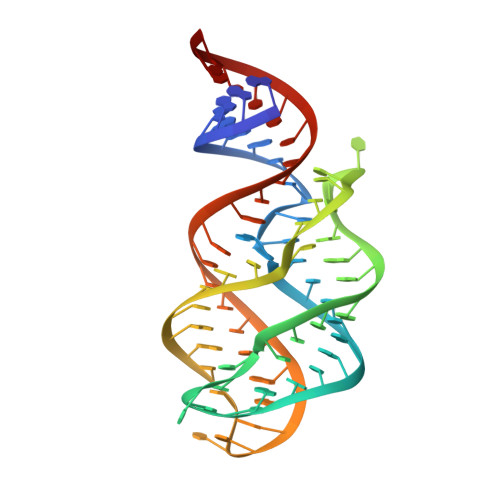
Synchronous RNA conformational changes trigger ordered phase transitions in crystals.
Ramakrishnan, S., Stagno, J.R., Conrad, C.E., Ding, J., Yu, P., Bhandari, Y.R., Lee, Y.T., Pauly, G., Yefanov, O., Wiedorn, M.O., Knoska, J., Oberthur, D., White, T.A., Barty, A., Mariani, V., Li, C., Brehm, W., Heinz, W.F., Magidson, V., Lockett, S., Hunter, M.S., Boutet, S., Zatsepin, N.A., Zuo, X., Grant, T.D., Pandey, S., Schmidt, M., Spence, J.C.H., Chapman, H.N., Wang, Y.X.(2021) Nat Commun 12: 1762-1762
- PubMed: 33741910 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-21838-5
- Primary Citation Related Structures:
6VWT, 6VWV - PubMed Abstract:
Time-resolved studies of biomacromolecular crystals have been limited to systems involving only minute conformational changes within the same lattice. Ligand-induced changes greater than several angstroms, however, are likely to result in solid-solid phase transitions, which require a detailed understanding of the mechanistic interplay between conformational and lattice transitions. Here we report the synchronous behavior of the adenine riboswitch aptamer RNA in crystal during ligand-triggered isothermal phase transitions. Direct visualization using polarized video microscopy and atomic force microscopy shows that the RNA molecules undergo cooperative rearrangements that maintain lattice order, whose cell parameters change distinctly as a function of time. The bulk lattice order throughout the transition is further supported by time-resolved diffraction data from crystals using an X-ray free electron laser. The synchronous molecular rearrangements in crystal provide the physical basis for studying large conformational changes using time-resolved crystallography and micro/nanocrystals.
- Structural Biophysics Laboratory, National Cancer Institute, Frederick, MD, USA.
Organizational Affiliation: