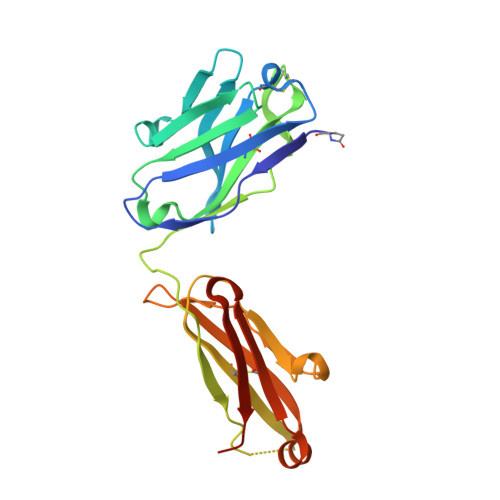
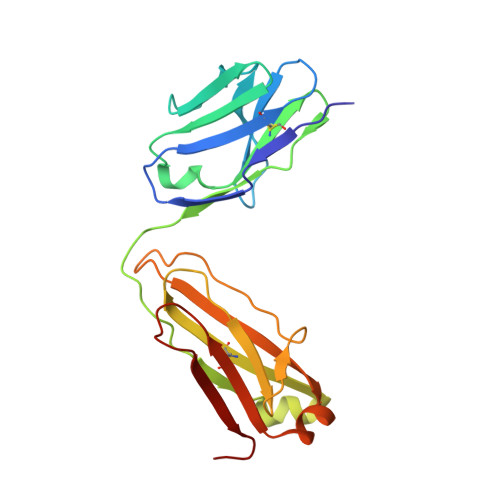
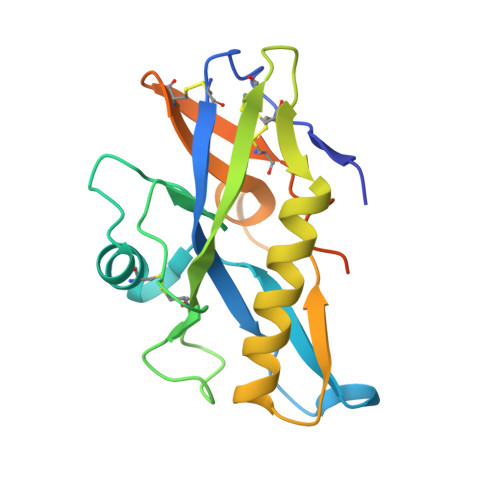
HIV-1 VRC01 Germline-Targeting Immunogens Select Distinct Epitope-Specific B Cell Receptors.
Lin, Y.R., Parks, K.R., Weidle, C., Naidu, A.S., Khechaduri, A., Riker, A.O., Takushi, B., Chun, J.H., Borst, A.J., Veesler, D., Stuart, A., Agrawal, P., Gray, M., Pancera, M., Huang, P.S., Stamatatos, L.(2020) Immunity 53: 840-851.e6
- PubMed: 33053332 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2020.09.007
- Primary Citation Related Structures:
6VLW - PubMed Abstract:
Activating precursor B cell receptors of HIV-1 broadly neutralizing antibodies requires specifically designed immunogens. Here, we compared the abilities of three such germline-targeting immunogens against the VRC01-class receptors to activate the targeted B cells in transgenic mice expressing the germline VH of the VRC01 antibody but diverse mouse light chains. Immunogen-specific VRC01-like B cells were isolated at different time points after immunization, their VH and VL genes were sequenced, and the corresponding antibodies characterized. VRC01 B cell sub-populations with distinct cross-reactivity properties were activated by each immunogen, and these differences correlated with distinct biophysical and biochemical features of the germline-targeting immunogens. Our study indicates that the design of effective immunogens to activate B cell receptors leading to protective HIV-1 antibodies will require a better understanding of how the biophysical properties of the epitope and its surrounding surface on the germline-targeting immunogen influence its interaction with the available receptor variants in vivo.
- Fred Hutchinson Cancer Research Center, Vaccines and Infectious Diseases Division, Seattle, WA, USA.
Organizational Affiliation: