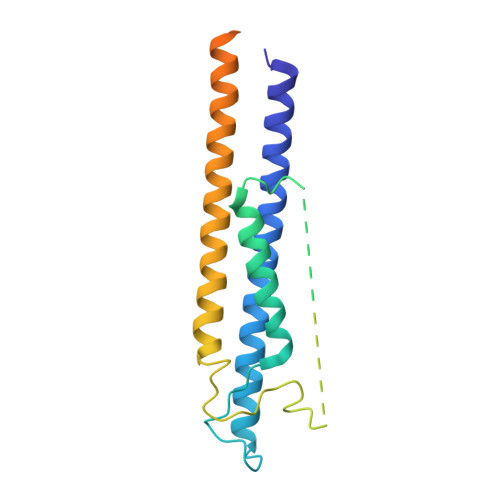
A non-canonical monovalent zinc finger stabilizes the integration of Cfp1 into the H3K4 methyltransferase complex COMPASS.
Yang, Y., Joshi, M., Takahashi, Y.H., Ning, Z., Qu, Q., Brunzelle, J.S., Skiniotis, G., Figeys, D., Shilatifard, A., Couture, J.F.(2020) Nucleic Acids Res 48: 421-431
- PubMed: 31724694 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz1037
- Primary Citation Related Structures:
6VHF - PubMed Abstract:
COMPlex ASsociating with SET1 (COMPASS) is a histone H3 Lys-4 methyltransferase that typically marks the promoter region of actively transcribed genes. COMPASS is a multi-subunit complex in which the catalytic unit, SET1, is required for H3K4 methylation. An important subunit known to regulate SET1 methyltransferase activity is the CxxC zinc finger protein 1 (Cfp1). Cfp1 binds to COMPASS and is critical to maintain high level of H3K4me3 in cells but the mechanisms underlying its stimulatory activity is poorly understood. In this study, we show that Cfp1 only modestly activates COMPASS methyltransferase activity in vitro. Binding of Cfp1 to COMPASS is in part mediated by a new type of monovalent zinc finger (ZnF). This ZnF interacts with the COMPASS's subunits RbBP5 and disruption of this interaction blunts its methyltransferase activity in cells and in vivo. Collectively, our studies reveal that a novel form of ZnF on Cfp1 enables its integration into COMPASS and contributes to epigenetic signaling.
- Shanghai Institute of Materia Medica-University of Ottawa Joint Research Centre on Systems and Personalized Pharmacology, University of Ottawa, Ottawa, ON K1H 8M5, Canada.
Organizational Affiliation: